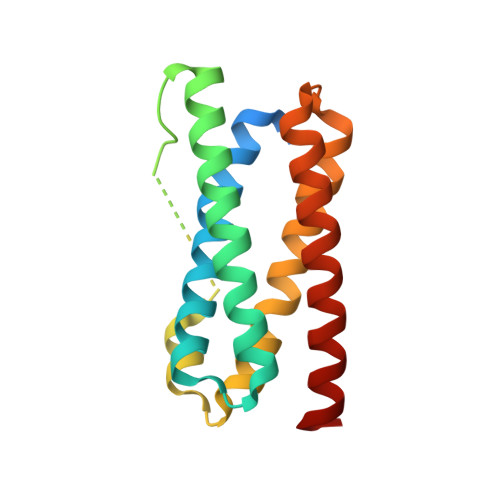
Structural basis for copper/silver binding by the Synechocystis metallochaperone CopM.
Zhao, S., Wang, X., Niu, G., Dong, W., Wang, J., Fang, Y., Lin, Y., Liu, L.(2016) Acta Crystallogr D Struct Biol 72: 997-1005
- PubMed: 27599732 Search on PubMed
- DOI: https://doi.org/10.1107/S2059798316011943
- Primary Citation Related Structures:
5FEJ, 5FFA, 5FFB, 5FFC, 5FFD, 5FFE - PubMed Abstract:
Copper homeostasis integrates multiple processes from sensing to storage and efflux out of the cell. CopM is a cyanobacterial metallochaperone, the gene for which is located upstream of a two-component system for copper resistance, but the molecular basis for copper recognition by this four-helical bundle protein is unknown. Here, crystal structures of CopM in apo, copper-bound and silver-bound forms are reported. Monovalent copper/silver ions are buried within the bundle core; divalent copper ions are found on the surface of the bundle. The monovalent copper/silver-binding site is constituted by two consecutive histidines and is conserved in a previously functionally unknown protein family. The structural analyses show two conformational states and suggest that flexibility in the first α-helix is related to the metallochaperone function. These results also reveal functional diversity from a protein family with a simple four-helical fold.
- Key Laboratory of Photobiology, CAS Center for Excellence in Molecular Plant Sciences, Institute of Botany, Chinese Academy of Sciences, Beijing 100093, People's Republic of China.
Organizational Affiliation: