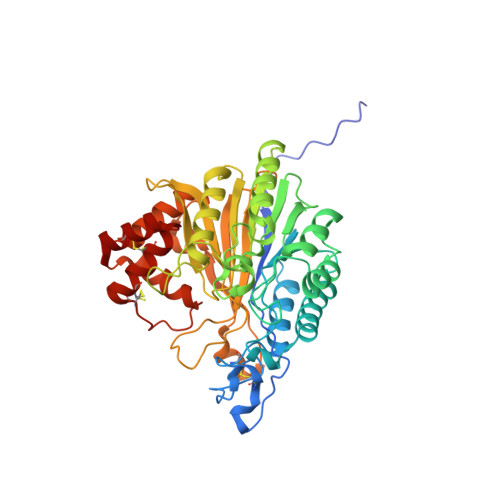
Structural Basis for Nucleotide Hydrolysis by the Acid Sphingomyelinase-like Phosphodiesterase SMPDL3A.
Gorelik, A., Illes, K., Superti-Furga, G., Nagar, B.(2016) J Biological Chem 291: 6376-6385
- PubMed: 26792860
- DOI: https://doi.org/10.1074/jbc.M115.711085
- Primary Citation of Related Structures:
5FC1, 5FC5, 5FC6, 5FC7, 5FCA, 5FCB - PubMed Abstract:
Sphingomyelin phosphodiesterase, acid-like 3A (SMPDL3A) is a member of a small family of proteins founded by the well characterized lysosomal enzyme, acid sphingomyelinase (ASMase). ASMase converts sphingomyelin into the signaling lipid, ceramide. It was recently discovered that, in contrast to ASMase, SMPDL3A is inactive against sphingomyelin and, surprisingly, can instead hydrolyze nucleoside diphosphates and triphosphates, which may play a role in purinergic signaling. As none of the ASMase-like proteins has been structurally characterized to date, the molecular basis for their substrate preferences is unknown. Here we report crystal structures of murine SMPDL3A, which represent the first structures of an ASMase-like protein. The catalytic domain consists of a central mixed β-sandwich surrounded by α-helices. Additionally, SMPDL3A possesses a unique C-terminal domain formed from a cluster of four α-helices that appears to distinguish this protein family from other phosphoesterases. We show that SMDPL3A is a di-zinc-dependent enzyme with an active site configuration that suggests a mechanism of phosphodiester hydrolysis by a metal-activated water molecule and protonation of the leaving group by a histidine residue. Co-crystal structures of SMPDL3A with AMP and α,β-methylene ADP (AMPCP) reveal that the substrate binding site accommodates nucleotides by establishing interactions with their base, sugar, and phosphate moieties, with the latter the major contributor to binding affinity. Our study provides the structural basis for SMPDL3A substrate specificity and sheds new light on the function of ASMase-like proteins.
- From the Department of Biochemistry and Groupe de Recherche Axe sur la Structure des Proteines, Faculty of Medicine, McGill University, Montreal, Quebec H3G 0B1, Canada and.
Organizational Affiliation: