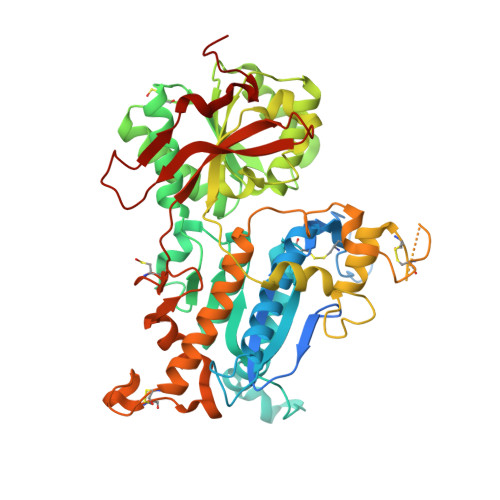
Structural basis for regulation of human calcium-sensing receptor by magnesium ions and an unexpected tryptophan derivative co-agonist.
Zhang, C., Zhang, T., Zou, J., Miller, C.L., Gorkhali, R., Yang, J.Y., Schilmiller, A., Wang, S., Huang, K., Brown, E.M., Moremen, K.W., Hu, J., Yang, J.J.(2016) Sci Adv 2: e1600241-e1600241
- PubMed: 27386547
- DOI: https://doi.org/10.1126/sciadv.1600241
- Primary Citation Related Structures:
5FBH, 5FBK - PubMed Abstract:
Ca(2+)-sensing receptors (CaSRs) modulate calcium and magnesium homeostasis and many (patho)physiological processes by responding to extracellular stimuli, including divalent cations and amino acids. We report the first crystal structure of the extracellular domain (ECD) of human CaSR bound with Mg(2+) and a tryptophan derivative ligand at 2.1 Å. The structure reveals key determinants for cooperative activation by metal ions and aromatic amino acids. The unexpected tryptophan derivative was bound in the hinge region between two globular ECD subdomains, and represents a novel high-affinity co-agonist of CaSR. The dissection of structure-function relations by mutagenesis, biochemical, and functional studies provides insights into the molecular basis of human diseases arising from CaSR mutations. The data also provide a novel paradigm for understanding the mechanism of CaSR-mediated signaling that is likely shared by the other family C GPCR [G protein (heterotrimeric guanine nucleotide-binding protein)-coupled receptor] members and can facilitate the development of novel CaSR-based therapeutics.
- Department of Chemistry, Center for Diagnostics and Therapeutics, Georgia State University, 50 Decatur Street, Atlanta, GA 30303, USA.
Organizational Affiliation: