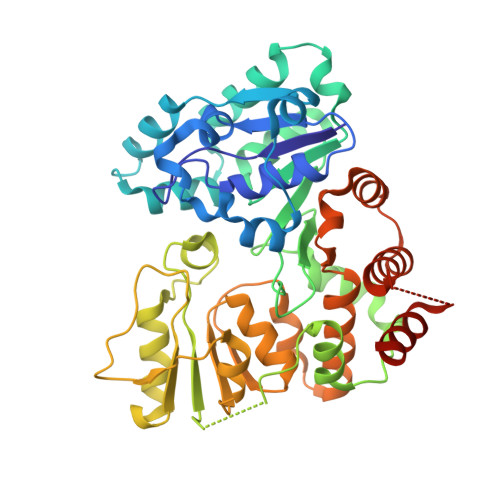
Bacterial beta-Kdo glycosyltransferases represent a new glycosyltransferase family (GT99).
Ovchinnikova, O.G., Mallette, E., Koizumi, A., Lowary, T.L., Kimber, M.S., Whitfield, C.(2016) Proc Natl Acad Sci U S A 113: E3120-E3129
- PubMed: 27199480
- DOI: https://doi.org/10.1073/pnas.1603146113
- Primary Citation Related Structures:
5FA0, 5FA1 - PubMed Abstract:
Kdo (3-deoxy-d-manno-oct-2-ulosonic acid) is an eight-carbon sugar mostly confined to Gram-negative bacteria. It is often involved in attaching surface polysaccharides to their lipid anchors. α-Kdo provides a bridge between lipid A and the core oligosaccharide in all bacterial LPSs, whereas an oligosaccharide of β-Kdo residues links "group 2" capsular polysaccharides to (lyso)phosphatidylglycerol. β-Kdo is also found in a small number of other bacterial polysaccharides. The structure and function of the prototypical cytidine monophosphate-Kdo-dependent α-Kdo glycosyltransferase from LPS assembly is well characterized. In contrast, the β-Kdo counterparts were not identified as glycosyltransferase enzymes by bioinformatics tools and were not represented among the 98 currently recognized glycosyltransferase families in the Carbohydrate-Active Enzymes database. We report the crystallographic structure and function of a prototype β-Kdo GT from WbbB, a modular protein participating in LPS O-antigen synthesis in Raoultella terrigena The β-Kdo GT has dual Rossmann-fold motifs typical of GT-B enzymes, but extensive deletions, insertions, and rearrangements result in a unique architecture that makes it a prototype for a new GT family (GT99). The cytidine monophosphate-binding site in the C-terminal α/β domain closely resembles the corresponding site in bacterial sialyltransferases, suggesting an evolutionary connection that is not immediately evident from the overall fold or sequence similarities.
- Department of Molecular and Cellular Biology, University of Guelph, Guelph, ON, Canada N1G 2W1;
Organizational Affiliation: