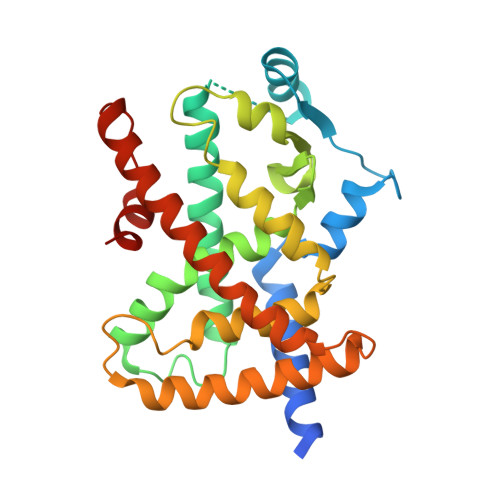
Screening of saponins and sapogenins from Medicago species as potential PPAR gamma agonists and X-ray structure of the complex PPAR gamma /caulophyllogenin.
Montanari, R., Capelli, D., Tava, A., Galli, A., Laghezza, A., Tortorella, P., Loiodice, F., Pochetti, G.(2016) Sci Rep 6: 27658-27658
- PubMed: 27283034
- DOI: https://doi.org/10.1038/srep27658
- Primary Citation of Related Structures:
5F9B - PubMed Abstract:
A series of saponins and sapogenins from Medicago species were tested for their ability to bind and activate the nuclear receptor PPARγ by SPR experiments and transactivation assay, respectively. The SPR analysis proved to be a very powerful and fast technique for screening a large number of compounds for their affinity to PPARγ and selecting the better candidates for further studies. Based on the obtained results, the sapogenin caulophyllogenin was proved to be a partial agonist towards PPARγ and the X-ray structure of its complex with PPARγ was also solved, in order to investigate the binding mode in the ligand binding domain of the nuclear receptor. This is the first known crystal structure of a sapogenin directly interacting with PPARγ. Another compound of the series, the echinocistic acid, showed antagonist activity towards PPARγ, a property that could be useful to inhibit the adipocyte differentiation which is a typical adverse effect of PPARγ agonists. This study confirms the interest on saponins and sapogenins as a valuable natural resource exploitable in the medical and food industry for ameliorating the metabolic syndrome.
- Istituto di Cristallografia, Consiglio Nazionale delle Ricerche, Montelibretti, 00015 Monterotondo Stazione, Roma, Italy.
Organizational Affiliation: