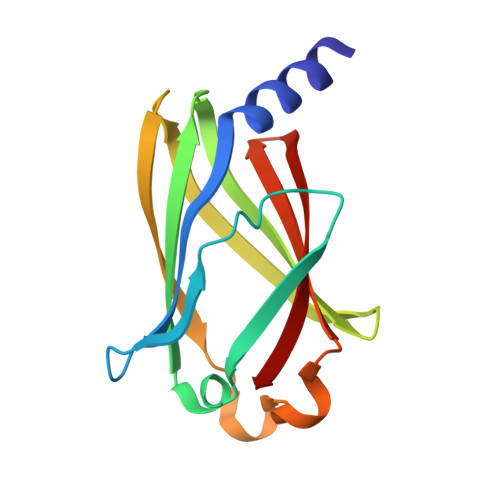
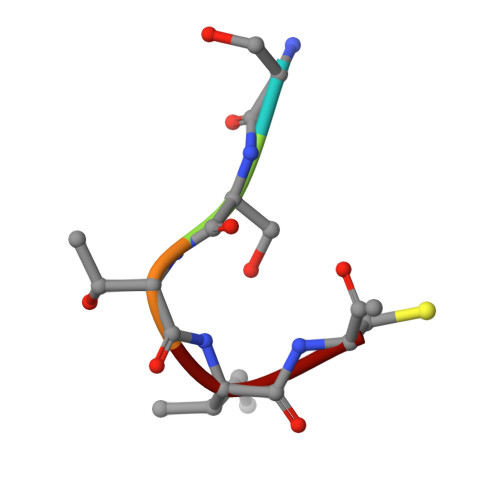
PDE6delta-mediated sorting of INPP5E into the cilium is determined by cargo-carrier affinity.
Fansa, E.K., Koesling, S.K., Zent, E., Wittinghofer, A., Ismail, S.(2016) Nat Commun 7: 11366
- PubMed: 27063844 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/ncomms11366
- Primary Citation Related Structures:
5F2U - PubMed Abstract:
The phosphodiesterase 6 delta subunit (PDE6δ) shuttles several farnesylated cargos between membranes. The cargo sorting mechanism between cilia and other compartments is not understood. Here we show using the inositol polyphosphate 5'-phosphatase E (INPP5E) and the GTP-binding protein (Rheb) that cargo sorting depends on the affinity towards PDE6δ and the specificity of cargo release. High-affinity cargo is exclusively released by the ciliary transport regulator Arl3, while low-affinity cargo is released by Arl3 and its non-ciliary homologue Arl2. Structures of PDE6δ/cargo complexes reveal the molecular basis of the sorting signal which depends on the residues at the -1 and -3 positions relative to farnesylated cysteine. Structure-guided mutation allows the generation of a low-affinity INPP5E mutant which loses exclusive ciliary localization. We postulate that the affinity to PDE6δ and the release by Arl2/3 in addition to a retention signal are the determinants for cargo sorting and enrichment at its destination.
- Max Planck Institute of Molecular Physiology, Otto-Hahn-Strasse 11, 44227 Dortmund, Germany.
Organizational Affiliation: