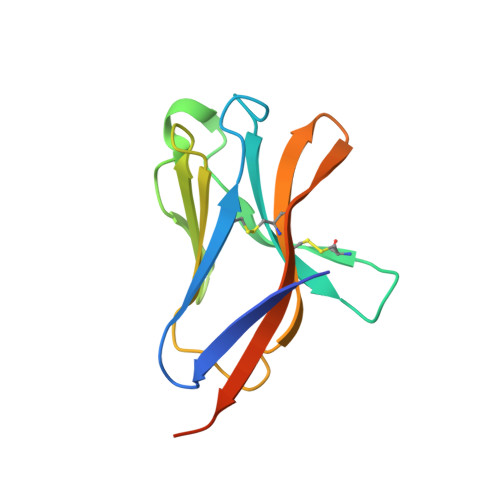
Neurodegenerative disease mutations in TREM2 reveal a functional surface and distinct loss-of-function mechanisms.
Kober, D.L., Alexander-Brett, J.M., Karch, C.M., Cruchaga, C., Colonna, M., Holtzman, M.J., Brett, T.J.(2016) Elife 5
- PubMed: 27995897 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.20391
- Primary Citation Related Structures:
5ELI - PubMed Abstract:
Genetic variations in the myeloid immune receptor TREM2 are linked to several neurodegenerative diseases. To determine how TREM2 variants contribute to these diseases, we performed structural and functional studies of wild-type and variant proteins. Our 3.1 Å TREM2 crystal structure revealed that mutations found in Nasu-Hakola disease are buried whereas Alzheimer's disease risk variants are found on the surface, suggesting that these mutations have distinct effects on TREM2 function. Biophysical and cellular methods indicate that Nasu-Hakola mutations impact protein stability and decrease folded TREM2 surface expression, whereas Alzheimer's risk variants impact binding to a TREM2 ligand. Additionally, the Alzheimer's risk variants appear to epitope map a functional surface on TREM2 that is unique within the larger TREM family. These findings provide a guide to structural and functional differences among genetic variants of TREM2, indicating that therapies targeting the TREM2 pathway should be tailored to these genetic and functional differences with patient-specific medicine approaches for neurodegenerative disorders.
- Molecular Microbiology and Microbial Pathogenesis Program, Washington University School of Medicine, St. Louis, United States.
Organizational Affiliation: