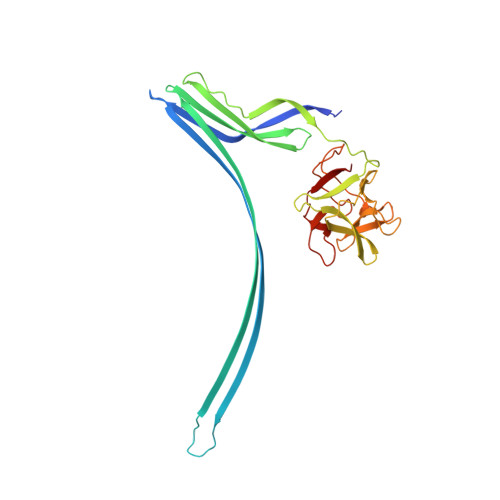
Crystal structure of an invertebrate cytolysin pore reveals unique properties and mechanism of assembly.
Podobnik, M., Savory, P., Rojko, N., Kisovec, M., Wood, N., Hambley, R., Pugh, J., Wallace, E.J., McNeill, L., Bruce, M., Liko, I., Allison, T.M., Mehmood, S., Yilmaz, N., Kobayashi, T., Gilbert, R.J., Robinson, C.V., Jayasinghe, L., Anderluh, G.(2016) Nat Commun 7: 11598-11598
- PubMed: 27176125 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/ncomms11598
- Primary Citation Related Structures:
5EC5 - PubMed Abstract:
The invertebrate cytolysin lysenin is a member of the aerolysin family of pore-forming toxins that includes many representatives from pathogenic bacteria. Here we report the crystal structure of the lysenin pore and provide insights into its assembly mechanism. The lysenin pore is assembled from nine monomers via dramatic reorganization of almost half of the monomeric subunit structure leading to a β-barrel pore ∼10 nm long and 1.6-2.5 nm wide. The lysenin pore is devoid of additional luminal compartments as commonly found in other toxin pores. Mutagenic analysis and atomic force microscopy imaging, together with these structural insights, suggest a mechanism for pore assembly for lysenin. These insights are relevant to the understanding of pore formation by other aerolysin-like pore-forming toxins, which often represent crucial virulence factors in bacteria.
- Department for Molecular Biology and Nanobiotechnology, National Institute of Chemistry, Hajdrihova 19, 1000 Ljubljana, Slovenia.
Organizational Affiliation: