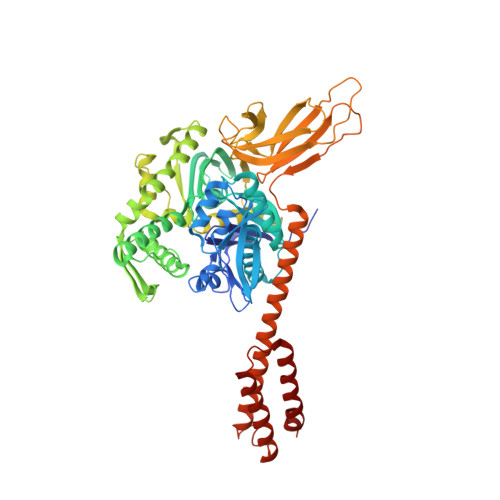
Close and Allosteric Opening of the Polypeptide-Binding Site in a Human Hsp70 Chaperone BiP.
Yang, J., Nune, M., Zong, Y., Zhou, L., Liu, Q.(2015) Structure 23: 2191-2203
- PubMed: 26655470 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2015.10.012
- Primary Citation Related Structures:
5E84, 5E85, 5E86 - PubMed Abstract:
Binding immunoglobulin protein (BiP), an essential and ubiquitous Hsp70 chaperone in the ER, plays a key role in protein folding and quality control. BiP contains two functional domains: a nucleotide-binding domain (NBD) and a substrate-binding domain (SBD). NBD binds and hydrolyzes ATP; the substrates for SBD are extended polypeptides. ATP binding allosterically accelerates polypeptide binding and release. Although crucial to the chaperone activity, the molecular mechanisms of polypeptide binding and allosteric coupling of BiP are poorly understood. Here, we present crystal structures of an intact human BiP in the ATP-bound state, the first intact eukaryotic Hsp70 structure, and isolated BiP-SBD with a peptide substrate bound representing the ADP-bound state. These structures and our biochemical analysis demonstrate that BiP has a unique NBD-SBD interface that is highly conserved only in eukaryotic Hsp70s found in the cytosol and ER to fortify its ATP-bound state and promote the opening of its polypeptide-binding pocket.
- Department of Physiology and Biophysics, Virginia Commonwealth University, Richmond, VA 23298, USA.
Organizational Affiliation: