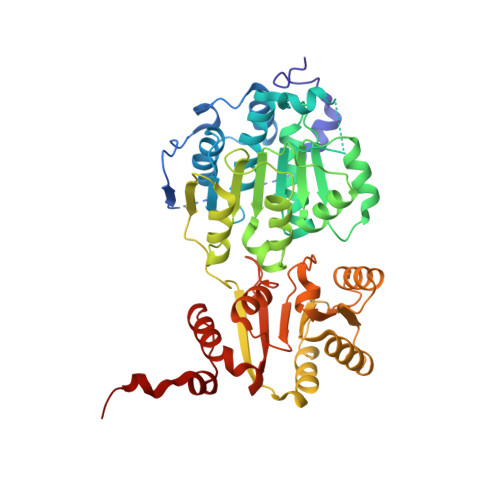
Autoinhibitory Interdomain Interactions and Subfamily-specific Extensions Redefine the Catalytic Core of the Human DEAD-box Protein DDX3.
Floor, S.N., Condon, K.J., Sharma, D., Jankowsky, E., Doudna, J.A.(2016) J Biological Chem 291: 2412-2421
- PubMed: 26598523 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M115.700625
- Primary Citation Related Structures:
5E7I, 5E7J, 5E7M - PubMed Abstract:
DEAD-box proteins utilize ATP to bind and remodel RNA and RNA-protein complexes. All DEAD-box proteins share a conserved core that consists of two RecA-like domains. The core is flanked by subfamily-specific extensions of idiosyncratic function. The Ded1/DDX3 subfamily of DEAD-box proteins is of particular interest as members function during protein translation, are essential for viability, and are frequently altered in human malignancies. Here, we define the function of the subfamily-specific extensions of the human DEAD-box protein DDX3. We describe the crystal structure of the subfamily-specific core of wild-type DDX3 at 2.2 Å resolution, alone and in the presence of AMP or nonhydrolyzable ATP. These structures illustrate a unique interdomain interaction between the two ATPase domains in which the C-terminal domain clashes with the RNA-binding surface. Destabilizing this interaction accelerates RNA duplex unwinding, suggesting that it is present in solution and inhibitory for catalysis. We use this core fragment of DDX3 to test the function of two recurrent medulloblastoma variants of DDX3 and find that both inactivate the protein in vitro and in vivo. Taken together, these results redefine the structural and functional core of the DDX3 subfamily of DEAD-box proteins.
- From the Department of Molecular and Cell Biology, Howard Hughes Medical Institute.
Organizational Affiliation: