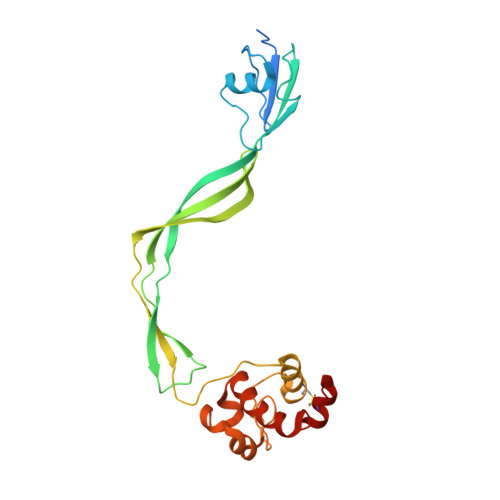
The structure of Resuscitation promoting factor B from M. tuberculosis reveals unexpected ubiquitin-like domains.
Ruggiero, A., Squeglia, F., Romano, M., Vitagliano, L., De Simone, A., Berisio, R.(2015) Biochim Biophys Acta 1860: 445-451
- PubMed: 26549874 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbagen.2015.11.001
- Primary Citation Related Structures:
5E27 - PubMed Abstract:
RpfB is a key factor in resuscitation from dormancy of Mycobacterium tuberculosis. This protein is a cell-wall glycosidase, which cleaves cell-wall peptidoglycan. RpfB is structurally complex and is composed of three types of domains, including a catalytic, a G5 and three DUF348 domains. Structural information is currently limited to a portion of the protein including only the catalytic and G5 domains. To gain insights into the structure and function of all domains we have undertaken structural investigations on a large protein fragment containing all three types of domains that constitute RpfB (RpfB3D). The structural features of RpfB3D have been investigated combining x-ray crystallography and biophysical studies. The crystal structure of RpfB3D provides the first structural characterization of a DUF348 domain and revealed an unexpected structural relationship with ubiquitin. The crystal structure also provides specific structural features of these domains explaining their frequent association with G5 domains. Results provided novel insights into the mechanism of peptidoglycan degradation necessary to the resuscitation of M. tuberculosis. Features of the DUF348 domain add structural data to a large set of proteins embedding this domain. Based on its structural similarity to ubiquitin and frequent association to the G5 domain, we propose to name this domain as G5-linked-Ubiquitin-like domain, UBLG5.
- Institute of Biostructures and Bioimaging, CNR, via Mezzocannone 16, Napoli, Italy.
Organizational Affiliation: