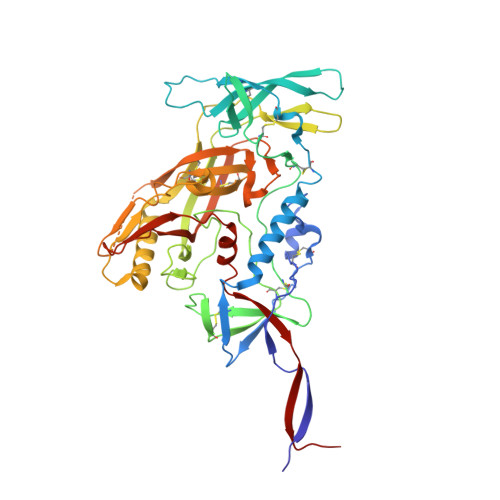
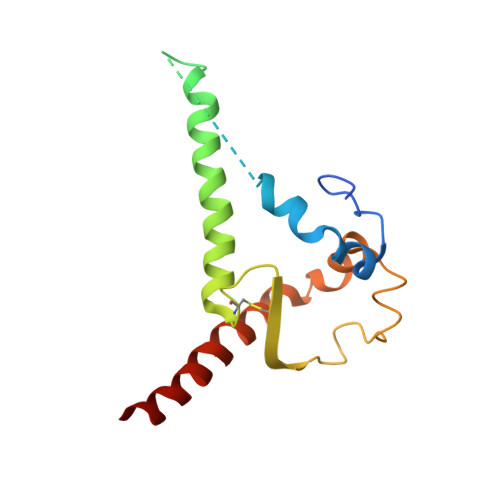
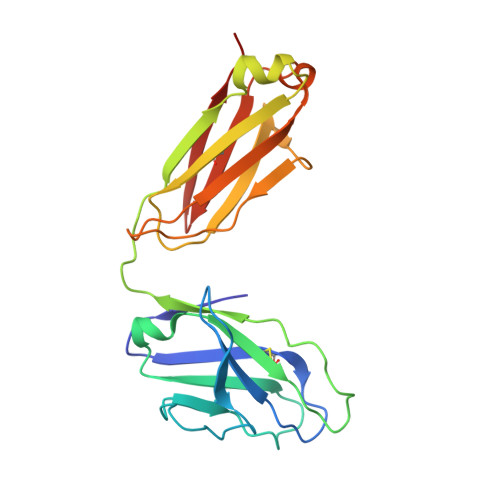
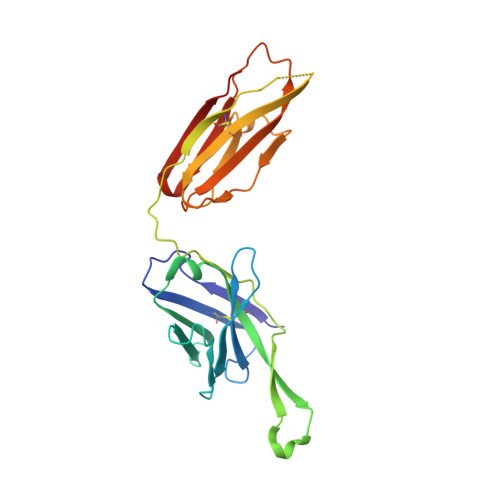
Minimally Mutated HIV-1 Broadly Neutralizing Antibodies to Guide Reductionist Vaccine Design.
Jardine, J.G., Sok, D., Julien, J.P., Briney, B., Sarkar, A., Liang, C.H., Scherer, E.A., Henry Dunand, C.J., Adachi, Y., Diwanji, D., Hsueh, J., Jones, M., Kalyuzhniy, O., Kubitz, M., Spencer, S., Pauthner, M., Saye-Francisco, K.L., Sesterhenn, F., Wilson, P.C., Galloway, D.M., Stanfield, R.L., Wilson, I.A., Burton, D.R., Schief, W.R.(2016) PLoS Pathog 12: e1005815-e1005815
- PubMed: 27560183 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.ppat.1005815
- Primary Citation Related Structures:
5D9Q, 5KZC - PubMed Abstract:
An optimal HIV vaccine should induce broadly neutralizing antibodies (bnAbs) that neutralize diverse viral strains and subtypes. However, potent bnAbs develop in only a small fraction of HIV-infected individuals, all contain rare features such as extensive mutation, insertions, deletions, and/or long complementarity-determining regions, and some are polyreactive, casting doubt on whether bnAbs to HIV can be reliably induced by vaccination. We engineered two potent VRC01-class bnAbs that minimized rare features. According to a quantitative features frequency analysis, the set of features for one of these minimally mutated bnAbs compared favorably with all 68 HIV bnAbs analyzed and was similar to antibodies elicited by common vaccines. This same minimally mutated bnAb lacked polyreactivity in four different assays. We then divided the minimal mutations into spatial clusters and dissected the epitope components interacting with those clusters, by mutational and crystallographic analyses coupled with neutralization assays. Finally, by synthesizing available data, we developed a working-concept boosting strategy to select the mutation clusters in a logical order following a germline-targeting prime. We have thus developed potent HIV bnAbs that may be more tractable vaccine goals compared to existing bnAbs, and we have proposed a strategy to elicit them. This reductionist approach to vaccine design, guided by antibody and antigen structure, could be applied to design candidate vaccines for other HIV bnAbs or protective Abs against other pathogens.
- Department of Immunology and Microbial Science, The Scripps Research Institute, La Jolla, California, United States of America.
Organizational Affiliation: