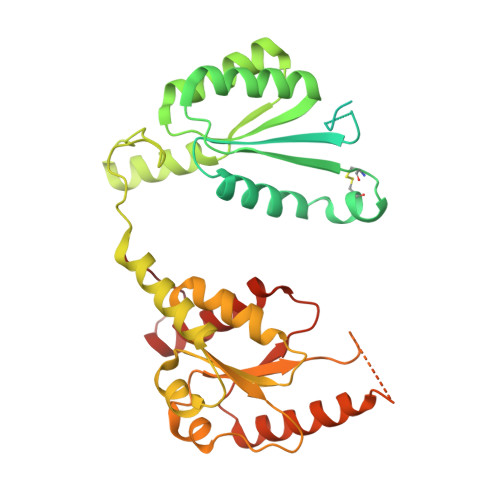
Structural Basis of Reversible Phosphorylation by Maize Pyruvate Orthophosphate Dikinase Regulatory Protein
Jiang, L., Chen, Y.B., Zheng, J., Chen, Z., Liu, Y., Tao, Y., Wu, W., Chen, Z., Wang, B.C.(2016) Plant Physiol 170: 732-741
- PubMed: 26620526 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1104/pp.15.01709
- Primary Citation Related Structures:
5D0N, 5D1F - PubMed Abstract:
Pyruvate orthophosphate dikinase (PPDK) is one of the most important enzymes in C4 photosynthesis. PPDK regulatory protein (PDRP) regulates the inorganic phosphate-dependent activation and ADP-dependent inactivation of PPDK by reversible phosphorylation. PDRP shares no significant sequence similarity with other protein kinases or phosphatases. To investigate the molecular mechanism by which PDRP carries out its dual and competing activities, we determined the crystal structure of PDRP from maize (Zea mays). PDRP forms a compact homo-dimer in which each protomer contains two separate N-terminal (NTD) and C-terminal (CTD) domains. The CTD includes several key elements for performing both phosphorylation and dephosphorylation activities: the phosphate binding loop (P-loop) for binding the ADP and inorganic phosphate substrates, residues Lys-274 and Lys-299 for neutralizing the negative charge, and residue Asp-277 for protonating and deprotonating the target Thr residue of PPDK to promote nucleophilic attack. Surprisingly, the NTD shares the same protein fold as the CTD and also includes a putative P-loop with AMP bound but lacking enzymatic activities. Structural analysis indicated that this loop may participate in the interaction with and regulation of PPDK. The NTD has conserved intramolecular and intermolecular disulfide bonds for PDRP dimerization. Moreover, PDRP is the first structure of the domain of unknown function 299 enzyme family reported. This study provides a structural basis for understanding the catalytic mechanism of PDRP and offers a foundation for the development of selective activators or inhibitors that may regulate photosynthesis.
- State Key Laboratory of Agrobiotechnology, China Agricultural University, Beijing 100193, China (L.J., J.Z., Zhenhang Chen, Y.L., W.W., Zhongzhou Chen);Photosynthesis Research Center, Key Laboratory of Photobiology, Institute of Botany, Chinese Academy of Sciences, Xiangshan, Beijing 100093, China (Y.-B.C., B.-C.W.); andBeijing Synchrotron Radiation Facility, Institute of High Energy Physics, Chinese Academy of Sciences, Beijing 100049, China (Y.T.).
Organizational Affiliation: