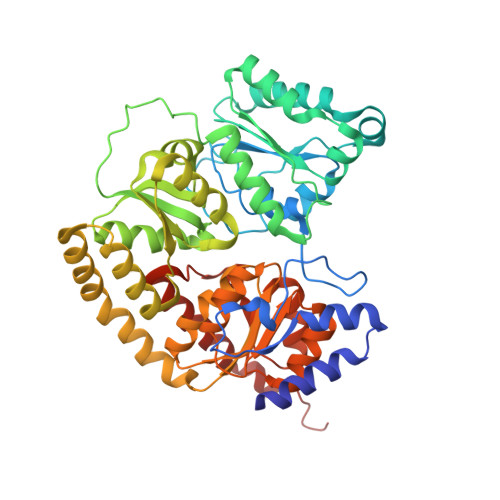
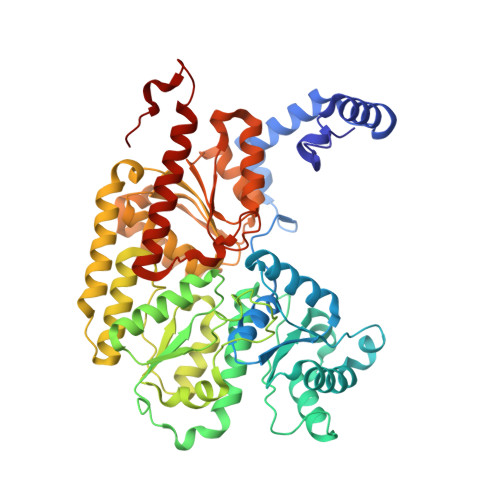
Evidence for Functionally Relevant Encounter Complexes in Nitrogenase Catalysis.
Owens, C.P., Katz, F.E., Carter, C.H., Luca, M.A., Tezcan, F.A.(2015) J Am Chem Soc 137: 12704-12712
- PubMed: 26360912 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/jacs.5b08310
- Primary Citation Related Structures:
5CX1 - PubMed Abstract:
Nitrogenase is the only enzyme that can convert atmospheric dinitrogen (N2) into biologically usable ammonia (NH3). To achieve this multielectron redox process, the nitrogenase component proteins, MoFe-protein (MoFeP) and Fe-protein (FeP), repeatedly associate and dissociate in an ATP-dependent manner, where one electron is transferred from FeP to MoFeP per association. Here, we provide experimental evidence that encounter complexes between FeP and MoFeP play a functional role in nitrogenase catalysis. The encounter complexes are stabilized by electrostatic interactions involving a positively charged patch on the β-subunit of MoFeP. Three single mutations (βAsn399Glu, βLys400Glu, and βArg401Glu) in this patch were generated in Azotobacter vinelandii MoFeP. All of the resulting variants displayed decreases in specific catalytic activity, with the βK400E mutation showing the largest effect. As simulated by the Thorneley-Lowe kinetic scheme, this single mutation lowered the rate constant for FeP-MoFeP association 5-fold. We also found that the βK400E mutation did not affect the coupling of ATP hydrolysis with electron transfer (ET) between FeP and MoFeP. These data suggest a mechanism where FeP initially forms encounter complexes on the MoFeP β-subunit surface en route to the ATP-activated, ET-competent complex over the αβ-interface.
- Department of Chemistry and Biochemistry, University of California, San Diego , La Jolla, California 92039, United States.
Organizational Affiliation: