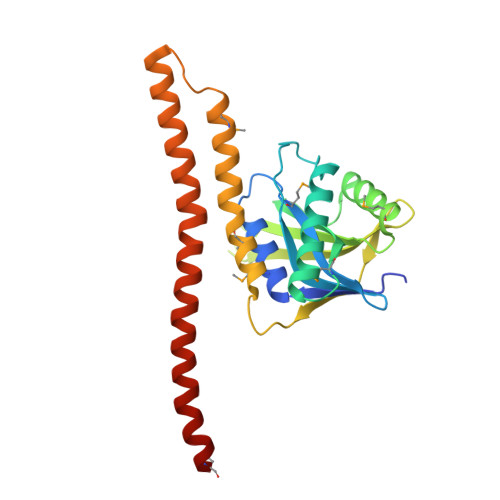
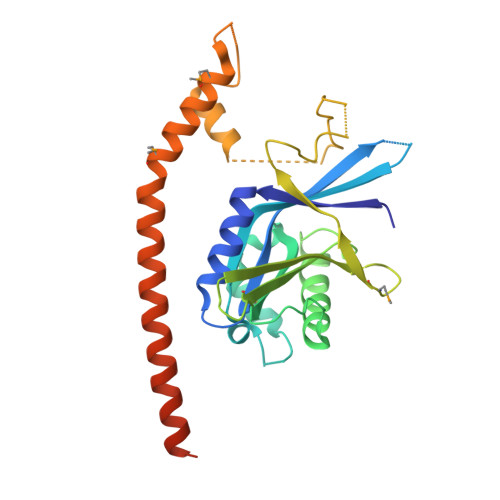
Higher-Order Assembly of BRCC36-KIAA0157 Is Required for DUB Activity and Biological Function.
Zeqiraj, E., Tian, L., Piggott, C.A., Pillon, M.C., Duffy, N.M., Ceccarelli, D.F., Keszei, A.F., Lorenzen, K., Kurinov, I., Orlicky, S., Gish, G.D., Heck, A.J., Guarne, A., Greenberg, R.A., Sicheri, F.(2015) Mol Cell 59: 970-983
- PubMed: 26344097 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.molcel.2015.07.028
- Primary Citation Related Structures:
5CW3, 5CW4, 5CW5, 5CW6 - PubMed Abstract:
BRCC36 is a Zn(2+)-dependent deubiquitinating enzyme (DUB) that hydrolyzes lysine-63-linked ubiquitin chains as part of distinct macromolecular complexes that participate in either interferon signaling or DNA-damage recognition. The MPN(+) domain protein BRCC36 associates with pseudo DUB MPN(-) proteins KIAA0157 or Abraxas, which are essential for BRCC36 enzymatic activity. To understand the basis for BRCC36 regulation, we have solved the structure of an active BRCC36-KIAA0157 heterodimer and an inactive BRCC36 homodimer. Structural and functional characterizations show how BRCC36 is switched to an active conformation by contacts with KIAA0157. Higher-order association of BRCC36 and KIAA0157 into a dimer of heterodimers (super dimers) was required for DUB activity and interaction with targeting proteins SHMT2 and RAP80. These data provide an explanation of how an inactive pseudo DUB allosterically activates a cognate DUB partner and implicates super dimerization as a new regulatory mechanism underlying BRCC36 DUB activity, subcellular localization, and biological function.
- Lunenfeld-Tanenbaum Research Institute, Mount Sinai Hospital, 600 University Avenue, Room 1090, Toronto, ON M5G 1X5, Canada.
Organizational Affiliation: