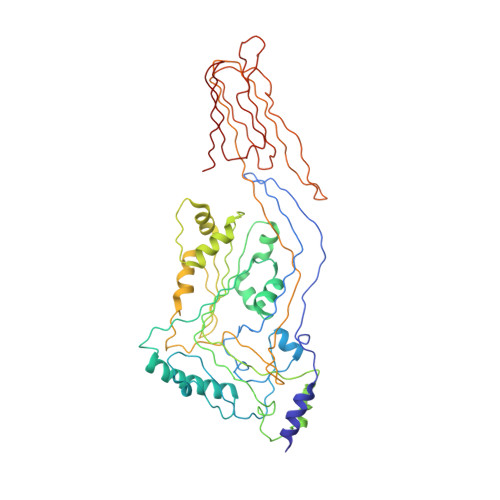
The Crystal Structure of Pneumolysin at 2.0 angstrom Resolution Reveals the Molecular Packing of the Pre-pore Complex.
Marshall, J.E., Faraj, B.H., Gingras, A.R., Lonnen, R., Sheikh, M.A., El-Mezgueldi, M., Moody, P.C., Andrew, P.W., Wallis, R.(2015) Sci Rep 5: 13293-13293
- PubMed: 26333773 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/srep13293
- Primary Citation Related Structures:
5CR6, 5CR8 - PubMed Abstract:
Pneumolysin is a cholesterol-dependent cytolysin (CDC) and virulence factor of Streptococcus pneumoniae. It kills cells by forming pores assembled from oligomeric rings in cholesterol-containing membranes. Cryo-EM has revealed the structures of the membrane-surface bound pre-pore and inserted-pore oligomers, however the molecular contacts that mediate these oligomers are unknown because high-resolution information is not available. Here we have determined the crystal structure of full-length pneumolysin at 1.98 Å resolution. In the structure, crystal contacts demonstrate the likely interactions that enable polymerisation on the cell membrane and the molecular packing of the pre-pore complex. The hemolytic activity is abrogated in mutants that disrupt these intermolecular contacts, highlighting their importance during pore formation. An additional crystal structure of the membrane-binding domain alone suggests that changes in the conformation of a tryptophan rich-loop at the base of the toxin promote monomer-monomer interactions upon membrane binding by creating new contacts. Notably, residues at the interface are conserved in other members of the CDC family, suggesting a common mechanism for pore and pre-pore assembly.
- Department of Infection, Immunity and Inflammation, University of Leicester, PO Box 138, Leicester, LE1 9HN UK.
Organizational Affiliation: