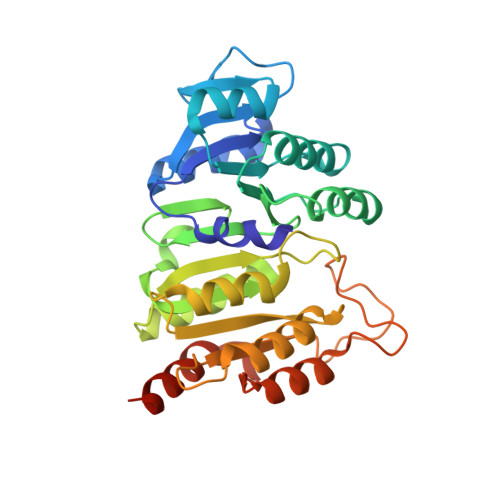
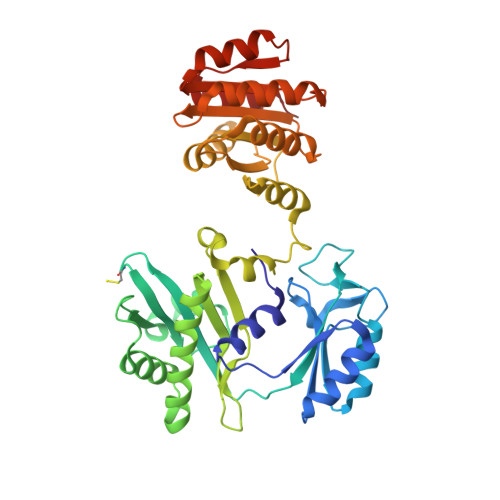
Structural basis for the binding of succinate to succinyl-CoA synthetase.
Huang, J., Fraser, M.E.(2016) Acta Crystallogr D Struct Biol 72: 912-921
- PubMed: 27487822 Search on PubMed
- DOI: https://doi.org/10.1107/S2059798316010044
- Primary Citation Related Structures:
5CAE - PubMed Abstract:
Succinyl-CoA synthetase catalyzes the only step in the citric acid cycle that provides substrate-level phosphorylation. Although the binding sites for the substrates CoA, phosphate, and the nucleotides ADP and ATP or GDP and GTP have been identified, the binding site for succinate has not. To determine this binding site, pig GTP-specific succinyl-CoA synthetase was crystallized in the presence of succinate, magnesium ions and CoA, and the structure of the complex was determined by X-ray crystallography to 2.2 Å resolution. Succinate binds in the carboxy-terminal domain of the β-subunit. The succinate-binding site is near both the active-site histidine residue that is phosphorylated in the reaction and the free thiol of CoA. The carboxy-terminal domain rearranges when succinate binds, burying this active site. However, succinate is not in position for transfer of the phosphoryl group from phosphohistidine. Here, it is proposed that when the active-site histidine residue has been phosphorylated by GTP, the phosphohistidine displaces phosphate and triggers the movement of the carboxylate of succinate into position to be phosphorylated. The structure shows why succinyl-CoA synthetase is specific for succinate and does not react appreciably with citrate nor with the other C4-dicarboxylic acids of the citric acid cycle, fumarate and oxaloacetate, but shows some activity with L-malate.
- Department of Biological Sciences, University of Calgary, 2500 University Drive NW, Calgary, Alberta T2N 1N4, Canada.
Organizational Affiliation: