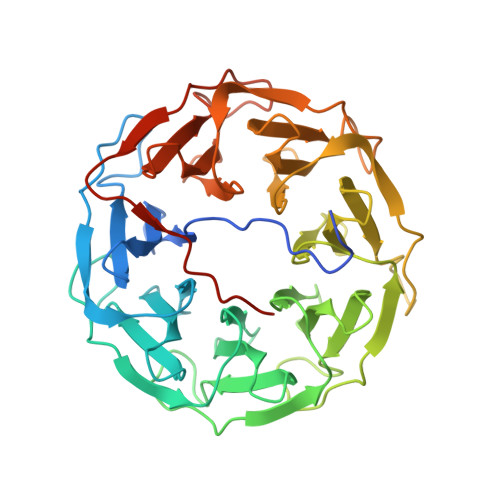
A Novel Fucose-binding Lectin from Photorhabdus luminescens (PLL) with an Unusual Heptabladed beta-Propeller Tetrameric Structure.
Kumar, A., Sykorova, P., Demo, G., Dobes, P., Hyrsl, P., Wimmerova, M.(2016) J Biological Chem 291: 25032-25049
- PubMed: 27758853
- DOI: https://doi.org/10.1074/jbc.M115.693473
- Primary Citation of Related Structures:
5C9L, 5C9O, 5C9P - PubMed Abstract:
Photorhabdus luminescens is known for its symbiosis with the entomopathogenic nematode Heterorhabditis bacteriophora and its pathogenicity toward insect larvae. A hypothetical protein from P. luminescens was identified, purified from the native source, and characterized as an l-fucose-binding lectin, named P. luminescens lectin (PLL). Glycan array and biochemical characterization data revealed PLL to be specific toward l-fucose and the disaccharide glycan 3,6-O-Me 2 -Glcβ1-4(2,3-O-Me 2 )Rhaα-O-(p-C 6 H 4 )-OCH 2 CH 2 NH 2 PLL was discovered to be a homotetramer with an intersubunit disulfide bridge. The crystal structures of native and recombinant PLL revealed a seven-bladed β-propeller fold creating seven putative fucose-binding sites per monomer. The crystal structure of the recombinant PLL·l-fucose complex confirmed that at least three sites were fucose-binding. Moreover, the crystal structures indicated that some of the other sites are masked either by the tetrameric nature of the lectin or by incorporation of the C terminus of the lectin into one of these sites. PLL exhibited an ability to bind to insect hemocytes and the cuticular surface of a nematode, H. bacteriophora.
- From the Central European Institute of Technology (CEITEC).
Organizational Affiliation: