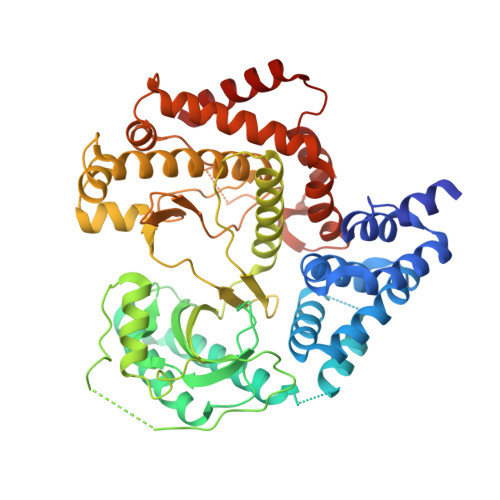
Using hydrogen deuterium exchange mass spectrometry to engineer optimized constructs for crystallization of protein complexes: Case study of PI4KIII beta with Rab11.
Fowler, M.L., McPhail, J.A., Jenkins, M.L., Masson, G.R., Rutaganira, F.U., Shokat, K.M., Williams, R.L., Burke, J.E.(2016) Protein Sci 25: 826-839
- PubMed: 26756197 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.2879
- Primary Citation Related Structures:
5C46, 5C4G - PubMed Abstract:
The ability of proteins to bind and interact with protein partners plays fundamental roles in many cellular contexts. X-ray crystallography has been a powerful approach to understand protein-protein interactions; however, a challenge in the crystallization of proteins and their complexes is the presence of intrinsically disordered regions. In this article, we describe an application of hydrogen deuterium exchange mass spectrometry (HDX-MS) to identify dynamic regions within type III phosphatidylinositol 4 kinase beta (PI4KIIIβ) in complex with the GTPase Rab11. This information was then used to design deletions that allowed for the production of diffraction quality crystals. Importantly, we also used HDX-MS to verify that the new construct was properly folded, consistent with it being catalytically and functionally active. Structures of PI4KIIIβ in an Apo state and bound to the potent inhibitor BQR695 in complex with both GTPγS and GDP loaded Rab11 were determined. This hybrid HDX-MS/crystallographic strategy revealed novel aspects of the PI4KIIIβ-Rab11 complex, as well as the molecular mechanism of potency of a PI4K specific inhibitor (BQR695). This approach is widely applicable to protein-protein complexes, and is an excellent strategy to optimize constructs for high-resolution structural approaches.
- Department of Biochemistry and Microbiology, University of Victoria, British Columbia, V8P 5C2, Canada.
Organizational Affiliation: