Mutational Constraints on Local Unfolding Inhibit the Rheological Adaptation of von Willebrand Factor.
Tischer, A., Campbell, J.C., Machha, V.R., Moon-Tasson, L., Benson, L.M., Sankaran, B., Kim, C., Auton, M.(2016) J Biological Chem 291: 3848-3859
- PubMed: 26677223 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M115.703850
- Primary Citation Related Structures:
5BV8 - PubMed Abstract:
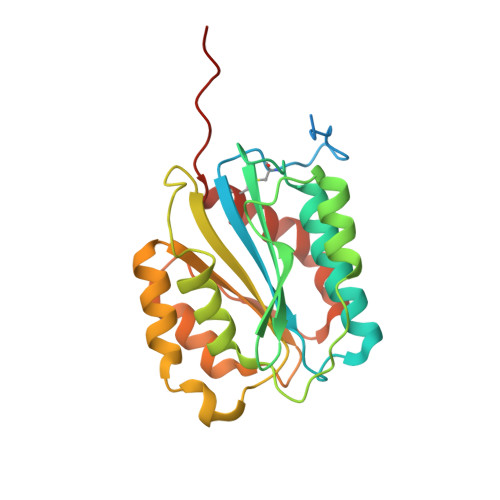
Unusually large von Willebrand factor (VWF), the first responder to vascular injury in primary hemostasis, is designed to capture platelets under the high shear stress of rheological blood flow. In type 2M von Willebrand disease, two rare mutations (G1324A and G1324S) within the platelet GPIbα binding interface of the VWF A1 domain impair the hemostatic function of VWF. We investigate structural and conformational effects of these mutations on the A1 domain's efficacy to bind collagen and adhere platelets under shear flow. These mutations enhance the thermodynamic stability, reduce the rate of unfolding, and enhance the A1 domain's resistance to limited proteolysis. Collagen binding affinity is not significantly affected indicating that the primary stabilizing effect of these mutations is to diminish the platelet binding efficiency under shear flow. The enhanced stability stems from the steric consequences of adding a side chain (G1324A) and additionally a hydrogen bond (G1324S) to His(1322) across the β2-β3 hairpin in the GPIbα binding interface, which restrains the conformational degrees of freedom and the overall flexibility of the native state. These studies reveal a novel rheological strategy in which the incorporation of a single glycine within the GPIbα binding interface of normal VWF enhances the probability of local unfolding that enables the A1 domain to conformationally adapt to shear flow while maintaining its overall native structure.
- From the Division of Hematology, Departments of Internal Medicine and Biochemistry and Molecular Biology, Mayo Clinic, Rochester, Minnesota 55905.
Organizational Affiliation: