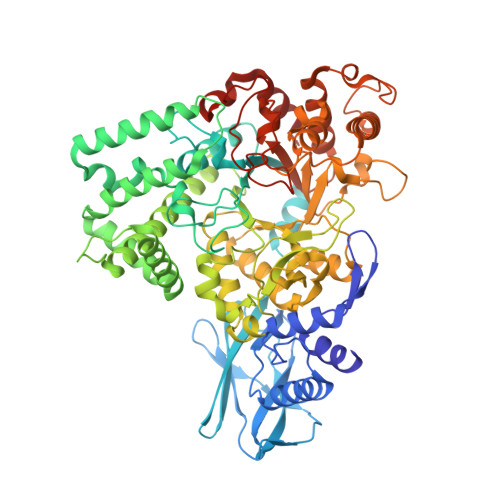
Crystal Structure of Amylomaltase from Corynebacterium glutamicum.
Joo, S., Kim, S., Seo, H., Kim, K.J.(2016) J Agric Food Chem 64: 5662-5670
- PubMed: 27366969 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jafc.6b02296
- Primary Citation Related Structures:
5B68 - PubMed Abstract:
Amylomaltase is an essential enzyme in maltose utilization and maltodextrin metabolism, and it has been industrially used for the production of cyclodextrin and modification of starch. We determined the crystal structure of amylomaltase from Corynebacterium glutamicum (CgAM) at a resolution of 1.7 Å. Although CgAM forms a dimer without NaCl, it exists as a monomer in physiological concentration of NaCl. CgAM is composed of N- and C-terminal domains, which can be further divided into two and four subdomains, respectively. It exhibits a unique structural feature at the functionally unknown N-domain and also shows two striking differences at the C-domain compared to other amylomaltases. These differences at extended edge of the substrate-binding site might affect substrate specificity for large cyclodextrin formation. The bis-tris methane and sulfate molecules bound at the substrate-binding site of our current structure mimic the binding of the hydroxyl groups of glucose bound at subsites -1 and -2, respectively.
- Structural and Molecular Biology Laboratory, School of Life Sciences and Biotechnology, Kyungpook National University , Daehak-ro 80, Buk-ku, Daegu 702-701, Republic of Korea.
Organizational Affiliation: