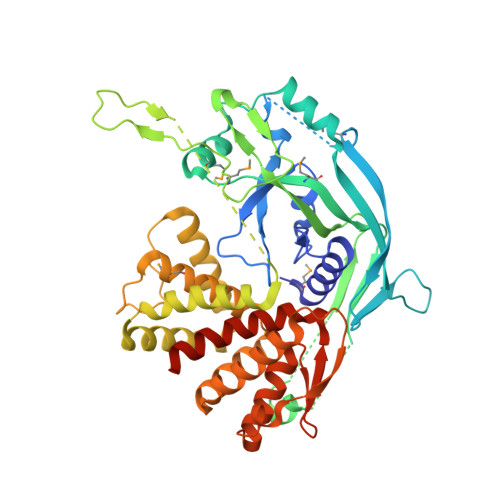
Pore-forming activity and structural autoinhibition of the gasdermin family.
Ding, J., Wang, K., Liu, W., She, Y., Sun, Q., Shi, J., Sun, H., Wang, D.C., Shao, F.(2016) Nature 535: 111-116
- PubMed: 27281216 Search on PubMed
- DOI: https://doi.org/10.1038/nature18590
- Primary Citation Related Structures:
5B5R - PubMed Abstract:
Inflammatory caspases cleave the gasdermin D (GSDMD) protein to trigger pyroptosis, a lytic form of cell death that is crucial for immune defences and diseases. GSDMD contains a functionally important gasdermin-N domain that is shared in the gasdermin family. The functional mechanism of action of gasdermin proteins is unknown. Here we show that the gasdermin-N domains of the gasdermin proteins GSDMD, GSDMA3 and GSDMA can bind membrane lipids, phosphoinositides and cardiolipin, and exhibit membrane-disrupting cytotoxicity in mammalian cells and artificially transformed bacteria. Gasdermin-N moved to the plasma membrane during pyroptosis. Purified gasdermin-N efficiently lysed phosphoinositide/cardiolipin-containing liposomes and formed pores on membranes made of artificial or natural phospholipid mixtures. Most gasdermin pores had an inner diameter of 10–14 nm and contained 16 symmetric protomers. The crystal structure of GSDMA3 showed an autoinhibited two-domain architecture that is conserved in the gasdermin family. Structure-guided mutagenesis demonstrated that the liposome-leakage and pore-forming activities of the gasdermin-N domain are required for pyroptosis. These findings reveal the mechanism for pyroptosis and provide insights into the roles of the gasdermin family in necrosis, immunity and diseases.