Crystal structure of the ectodomain from a LDLR close homologue in complex with its physiological ligand.
Hirai, H., Yasui, N., Yamashita, K., Tabata, S., Yamamoto, M., Takagi, J., Nogi, T.To be published.
Experimental Data Snapshot
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Reelin | 725 | Mus musculus | Mutation(s): 1 Gene Names: Reln, Rl EC: 3.4.21 | 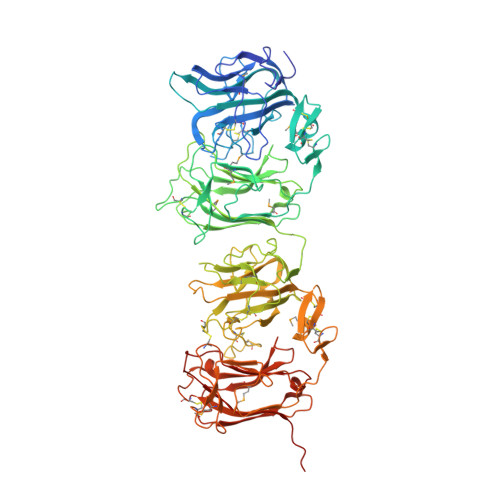 | |
UniProt & NIH Common Fund Data Resources | |||||
IMPC: MGI:103022 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q60841 | ||||
Glycosylation | |||||
| Glycosylation Sites: 1 | Go to GlyGen: Q60841-1 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Low density lipoprotein receptor-related protein 8, apolipoprotein e receptor, isoform CRA_e | 570 | Homo sapiens | Mutation(s): 0 Gene Names: LRP8, hCG_33395 | 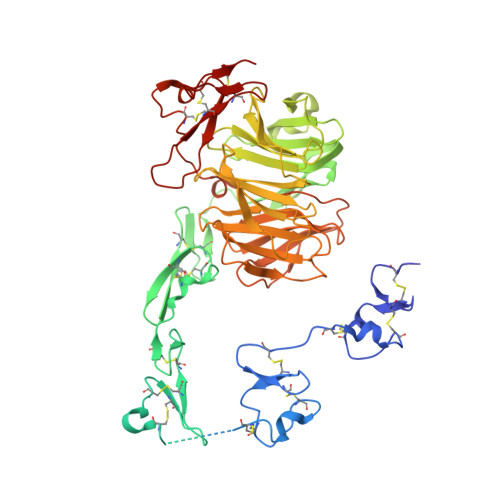 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: Q14114 GTEx: ENSG00000157193 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q14114 | ||||
Glycosylation | |||||
| Glycosylation Sites: 1 | Go to GlyGen: Q14114-1 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 2 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| NAG Download:Ideal Coordinates CCD File | I [auth A], J [auth A], O [auth B], T [auth C], Y [auth D] | 2-acetamido-2-deoxy-beta-D-glucopyranose C8 H15 N O6 OVRNDRQMDRJTHS-FMDGEEDCSA-N |  | ||
| CA Download:Ideal Coordinates CCD File | E [auth A] F [auth A] G [auth A] H [auth A] K [auth B] | CALCIUM ION Ca BHPQYMZQTOCNFJ-UHFFFAOYSA-N |  | ||
| Modified Residues 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Type | Formula | 2D Diagram | Parent |
| MSE Query on MSE | A, C | L-PEPTIDE LINKING | C5 H11 N O2 Se |  | MET |
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 205.951 | α = 90 |
| b = 205.951 | β = 90 |
| c = 169.84 | γ = 120 |
| Software Name | Purpose |
|---|---|
| REFMAC | refinement |
| XDS | data reduction |
| XSCALE | data scaling |
| autoSHARP | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| JSPS | Japan | 22247010 |
| JSPS | Japan | 20770084 |
| JSPS | Japan | 05J09821 |
| MEXT | Japan | 17082004 |