Crystal structure of mouse SAHH complexed with noraristeromycin
Kusakabe, Y., Ishihara, M., Tanaka, N.To be published.
Experimental Data Snapshot
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Adenosylhomocysteinase | A, B [auth C] | 432 | Mus musculus | Mutation(s): 0 Gene Names: Ahcy EC: 3.3.1.1 (PDB Primary Data), 3.13.2.1 (UniProt) | 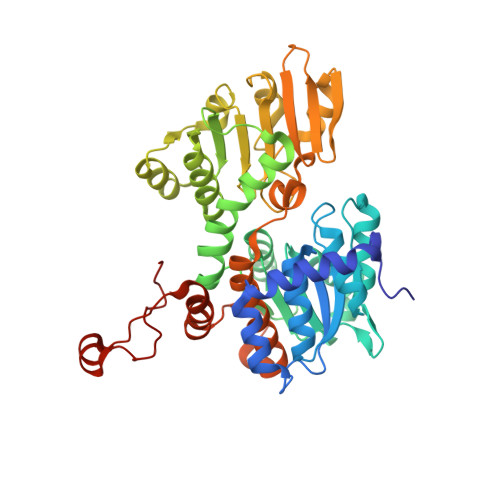 |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for P50247 (Mus musculus) Explore P50247 Go to UniProtKB: P50247 | |||||
IMPC: MGI:87968 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P50247 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 3 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| NAD Query on NAD | C [auth A], F [auth C] | NICOTINAMIDE-ADENINE-DINUCLEOTIDE C21 H27 N7 O14 P2 BAWFJGJZGIEFAR-NNYOXOHSSA-N |  | ||
| NRN Query on NRN | D [auth A], G [auth C] | (1S,2R,3S,4R)-4-(6-aminopurin-9-yl)cyclopentane-1,2,3-triol C10 H13 N5 O3 VFKHECGAEJNAMV-HETMPLHPSA-N |  | ||
| NA Query on NA | E [auth A], H [auth C] | SODIUM ION Na FKNQFGJONOIPTF-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 100.442 | α = 90 |
| b = 104.459 | β = 90 |
| c = 176.744 | γ = 90 |
| Software Name | Purpose |
|---|---|
| REFMAC | refinement |