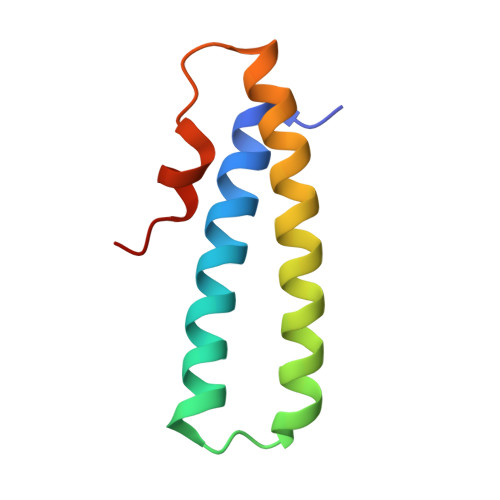
The Structure of the Toxin and Type Six Secretion System Substrate Tse2 in Complex with its Immunity Protein.
Robb, C.S., Robb, M., Nano, F.E., Boraston, A.B.(2016) Structure 24: 277
- PubMed: 26749446
- DOI: https://doi.org/10.1016/j.str.2015.11.012
- Primary Citation Related Structures:
5AKO - PubMed Abstract:
Tse2 is a cytoactive toxin secreted by a type six secretion apparatus of Pseudomonas aeruginosa. The Tse2 toxin naturally attacks a target in the cytoplasm of bacterial cells, and can cause toxicity if artificially introduced into eukaryotic cells. The X-ray crystal structure of the complex of Tse2 and its cognate immunity protein Tsi2 revealed a heterotetrameric structure with an extensive binding interface. Structural identity was found between Tse2 and NAD-dependent enzymes, especially ADP-ribosylating toxins, which facilitated the identification of the Tse2 active site and revealed it to be occluded upon binding the inhibitor Tsi2. The structural identity shared with NAD-dependent enzymes, including conserved catalytic residues, suggests that the mechanism of Tse2 toxicity may be NAD dependent.
- Department of Biochemistry and Microbiology, University of Victoria, Victoria, BC V8W 3P6, Canada.
Organizational Affiliation: