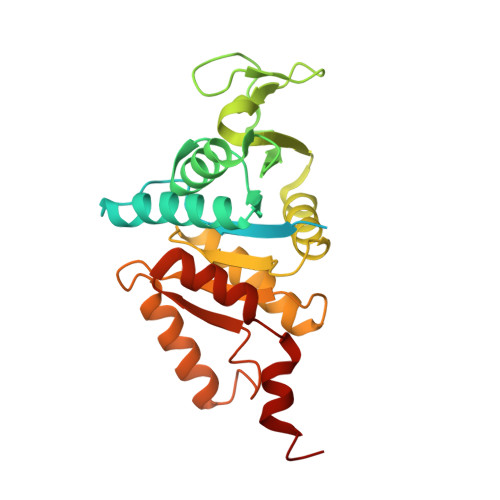
The Three-Dimensional Structure of "Lonely Guy" from Claviceps Purpurea Provides Insights Into the Phosphoribohydrolase Function of Rossmann Fold-Containing Lysine Decarboxylase-Like Proteins.
Dzurova, L., Forneris, F., Savino, S., Galuszka, P., Vrabka, J., Frebort, I.(2015) Proteins 83: 1539
- PubMed: 26010010
- DOI: https://doi.org/10.1002/prot.24835
- Primary Citation of Related Structures:
5AJT, 5AJU - PubMed Abstract:
The recently discovered cytokinin (CK)-specific phosphoribohydrolase "Lonely Guy" (LOG) is a key enzyme of CK biosynthesis, converting inactive CK nucleotides into biologically active free bases. We have determined the crystal structures of LOG from Claviceps purpurea (cpLOG) and its complex with the enzymatic product phosphoribose. The structures reveal a dimeric arrangement of Rossmann folds, with the ligands bound to large pockets at the interface between cpLOG monomers. Structural comparisons highlight the homology of cpLOG to putative lysine decarboxylases. Extended sequence analysis enabled identification of a distinguishing LOG sequence signature. Taken together, our data suggest phosphoribohydrolase activity for several proteins of unknown function.
- Centre of the Region Haná for Biotechnological and Agricultural Research, Faculty of Science, Palacký University, Olomouc, 783 71, Czech Republic.
Organizational Affiliation: