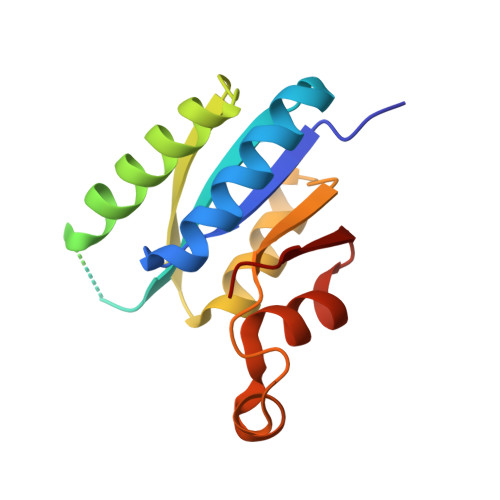
A Universal Stress Protein (Usp) in Mycobacteria Binds Camp
Banerjee, A., Adolph, R.S., Gopalakrishnapai, J., Kleinboelting, S., Emmerich, C., Steegborn, C., Visweswariah, S.S.(2015) J Biological Chem 290: 12731
- PubMed: 25802331
- DOI: https://doi.org/10.1074/jbc.M115.644856
- Primary Citation of Related Structures:
5AHW - PubMed Abstract:
Mycobacteria are endowed with rich and diverse machinery for the synthesis, utilization, and degradation of cAMP. The actions of cyclic nucleotides are generally mediated by binding of cAMP to conserved and well characterized cyclic nucleotide binding domains or structurally distinct cGMP-specific and -regulated cyclic nucleotide phosphodiesterase, adenylyl cyclase, and E. coli transcription factor FhlA (GAF) domain-containing proteins. Proteins with cyclic nucleotide binding and GAF domains can be identified in the genome of mycobacterial species, and some of them have been characterized. Here, we show that a significant fraction of intracellular cAMP is bound to protein in mycobacterial species, and by using affinity chromatography techniques, we identify specific universal stress proteins (USP) as abundantly expressed cAMP-binding proteins in slow growing as well as fast growing mycobacteria. We have characterized the biochemical and thermodynamic parameters for binding of cAMP, and we show that these USPs bind cAMP with a higher affinity than ATP, an established ligand for other USPs. We determined the structure of the USP MSMEG_3811 bound to cAMP, and we confirmed through structure-guided mutagenesis, the residues important for cAMP binding. This family of USPs is conserved in all mycobacteria, and we suggest that they serve as "sinks" for cAMP, making this second messenger available for downstream effectors as and when ATP levels are altered in the cell.
- From the Department of Molecular Reproduction, Development and Genetics, Indian Institute of Science, Bangalore 560012, India and.
Organizational Affiliation: