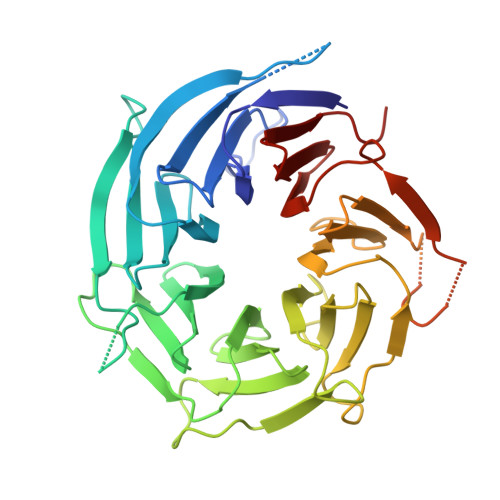
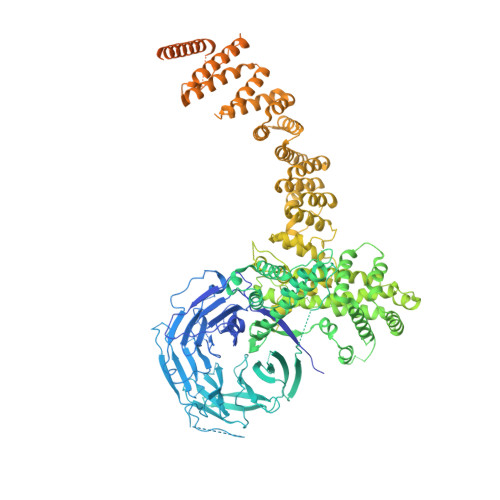
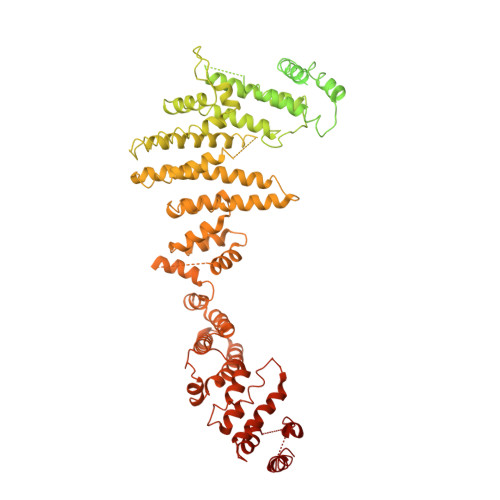
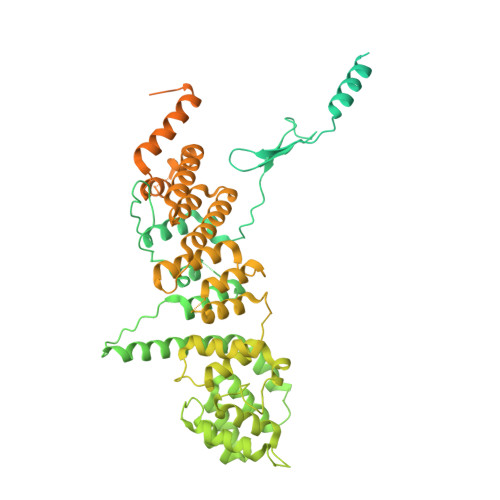
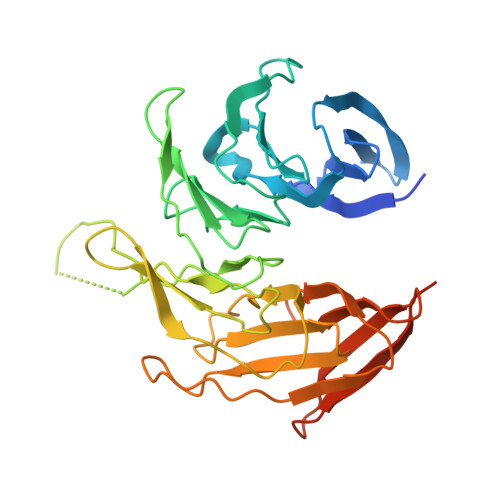
In Situ Structural Analysis of the Human Nuclear Pore Complex
Von Appen, A., Kosinski, J., Sparks, L., Ori, A., Diguilio, A., Vollmer, B., Mackmull, M., Banterle, N., Parca, L., Kastritis, P., Buczak, K., Mosalaganti, S., Hagen, W., Andres-Pons, A., Lemke, E.A., Bork, P., Antonin, W., Glavy, J.S., Bui, K.H., Beck, M.(2015) Nature 526: 140
- PubMed: 26416747 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature15381
- Primary Citation Related Structures:
5A9Q - PubMed Abstract:
Nuclear pore complexes are fundamental components of all eukaryotic cells that mediate nucleocytoplasmic exchange. Determining their 110-megadalton structure imposes a formidable challenge and requires in situ structural biology approaches. Of approximately 30 nucleoporins (Nups), 15 are structured and form the Y and inner-ring complexes. These two major scaffolding modules assemble in multiple copies into an eight-fold rotationally symmetric structure that fuses the inner and outer nuclear membranes to form a central channel of ~60 nm in diameter. The scaffold is decorated with transport-channel Nups that often contain phenylalanine-repeat sequences and mediate the interaction with cargo complexes. Although the architectural arrangement of parts of the Y complex has been elucidated, it is unclear how exactly it oligomerizes in situ. Here we combine cryo-electron tomography with mass spectrometry, biochemical analysis, perturbation experiments and structural modelling to generate, to our knowledge, the most comprehensive architectural model of the human nuclear pore complex to date. Our data suggest previously unknown protein interfaces across Y complexes and to inner-ring complex members. We show that the transport-channel Nup358 (also known as Ranbp2) has a previously unanticipated role in Y-complex oligomerization. Our findings blur the established boundaries between scaffold and transport-channel Nups. We conclude that, similar to coated vesicles, several copies of the same structural building block--although compositionally identical--engage in different local sets of interactions and conformations.
- European Molecular Biology Laboratory, Structural and Computational Biology Unit, Heidelberg, Germany.
Organizational Affiliation: