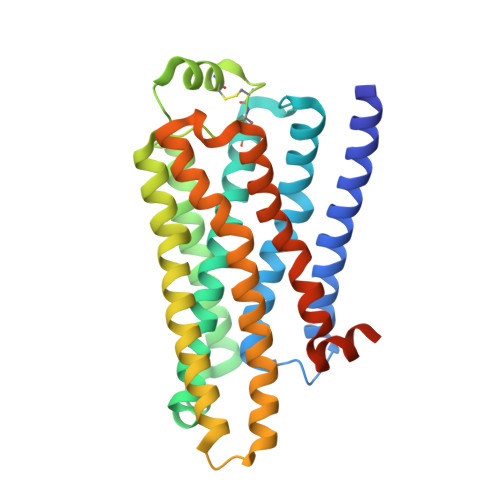
Pharmacological Analysis and Structure Determination of 7-Methylcyanopindolol-Bound Beta1-Adrenergic Receptor.
Sato, T., Baker, J., Warne, T., Brown, G., Leslie, A., Congreve, M., Tate, C.(2015) Mol Pharmacol 88: 1024
- PubMed: 26385885
- DOI: https://doi.org/10.1124/mol.115.101030
- Primary Citation of Related Structures:
5A8E - PubMed Abstract:
Comparisons between structures of the β1-adrenergic receptor (AR) bound to either agonists, partial agonists, or weak partial agonists led to the proposal that rotamer changes of Ser(5.46), coupled to a contraction of the binding pocket, are sufficient to increase the probability of receptor activation. (RS)-4-[3-(tert-butylamino)-2-hydroxypropoxy]-1H-indole-2-carbonitrile (cyanopindolol) is a weak partial agonist of β1AR and, based on the hypothesis above, we predicted that the addition of a methyl group to form 4-[(2S)-3-(tert-butylamino)-2-hydroxypropoxy]-7-methyl-1H-indole-2-carbonitrile (7-methylcyanopindolol) would dramatically reduce its efficacy. An eight-step synthesis of 7-methylcyanopindolol was developed and its pharmacology was analyzed. 7-Methylcyanopindolol bound with similar affinity to cyanopindolol to both β1AR and β2AR. As predicted, the efficacy of 7-methylcyanopindolol was reduced significantly compared with cyanopindolol, acting as a very weak partial agonist of turkey β1AR and an inverse agonist of human β2AR. The structure of 7-methylcyanopindolol-bound β1AR was determined to 2.4-Å resolution and found to be virtually identical to the structure of cyanopindolol-bound β1AR. The major differences in the orthosteric binding pocket are that it has expanded by 0.3 Å in 7-methylcyanopindolol-bound β1AR and the hydroxyl group of Ser(5.46) is positioned 0.8 Å further from the ligand, with respect to the position of the Ser(5.46) side chain in cyanopindolol-bound β1AR. Thus, the molecular basis for the reduction in efficacy of 7-methylcyanopindolol compared with cyanopindolol may be regarded as the opposite of the mechanism proposed for the increase in efficacy of agonists compared with antagonists.
- MRC Laboratory of Molecular Biology, Cambridge Biomedical Campus, Cambridge, United Kingdom (T.S., T.W., A.G.W.L., C.G.T.); Heptares Therapeutics Ltd, Welwyn Garden City, United Kingdom (G.A.B., M.C.); School of Life Sciences, University of Nottingham, Medical School, Queen's Medical Centre, Nottingham, United Kingdom (J.B.); KEK High Energy Accelerator Research Organization, Institute of Materials Structure Science, Structural Biology Research Center, Tsukuba, Japan (T.S.).
Organizational Affiliation: