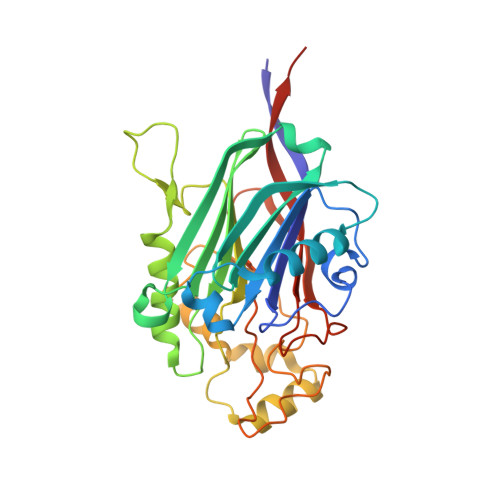
Crystal Structures of Type-II Inositol Polyphosphate 5-Phosphatase Inpp5B with Synthetic Inositol Polyphosphate Surrogates Reveal New Mechanistic Insights for the Inositol 5-Phosphatase Family.
Mills, S.J., Silvander, C., Cozier, G., Tresaugues, L., Nordlund, P., Potter, B.V.L.(2016) Biochemistry 55: 1384
- PubMed: 26854536 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.biochem.5b00838
- Primary Citation Related Structures:
5A7I, 5A7J - PubMed Abstract:
The inositol polyphosphate 5-phosphatase INPP5B hydrolyzes the 5-phosphate group from water- and lipid-soluble signaling messengers. Two synthetic benzene and biphenyl polyphosphates (BzP/BiPhPs), simplified surrogates of inositol phosphates and phospholipid headgroups, were identified by thermodynamic studies as potent INPP5B ligands. The X-ray structure of the complex between INPP5B and biphenyl 3,3',4,4',5,5'-hexakisphosphate [BiPh(3,3',4,4',5,5')P6, IC50 5.5 μM] was determined at 2.89 Å resolution. One inhibitor pole locates in the phospholipid headgroup binding site and the second solvent-exposed ring binds to the His-Tag of another INPP5B molecule, while a molecule of inorganic phosphate is also present in the active site. Benzene 1,2,3-trisphosphate [Bz(1,2,3)P3] [one ring of BiPh(3,3',4,4',5,5')P6] inhibits INPP5B ca. 6-fold less potently. Co-crystallization with benzene 1,2,4,5-tetrakisphosphate [Bz(1,2,4,5)P4, IC50 = 6.3 μM] yielded a structure refined at 2.9 Å resolution. Conserved residues among the 5-phosphatase family mediate interactions with Bz(1,2,4,5)P4 and BiPh(3,3',4,4',5,5')P6 similar to those with the polar groups present in positions 1, 4, 5, and 6 on the inositol ring of the substrate. 5-Phosphatase specificity most likely resides in the variable zone located close to the 2- and 3-positions of the inositol ring, offering insights to inhibitor design. We propose that the inorganic phosphate present in the INPP5B-BiPh(3,3',4,4',5,5')P6 complex mimics the postcleavage substrate 5-phosphate released by INPP5B in the catalytic site, allowing elucidation of two new key features in the catalytic mechanism proposed for the family of phosphoinositide 5-phosphatases: first, the involvement of the conserved Arg-451 in the interaction with the 5-phosphate and second, identification of the water molecule that initiates 5-phosphate hydrolysis. Our model also has implications for the proposed "moving metal" mechanism.
- Wolfson Laboratory of Medicinal Chemistry, Department of Pharmacy and Pharmacology, University of Bath , Bath BA2 7AY, U.K.
Organizational Affiliation: