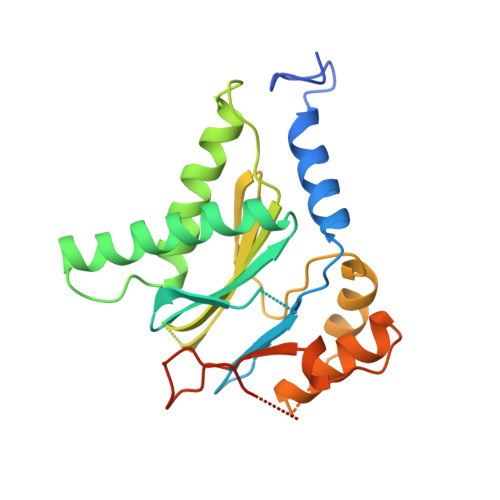
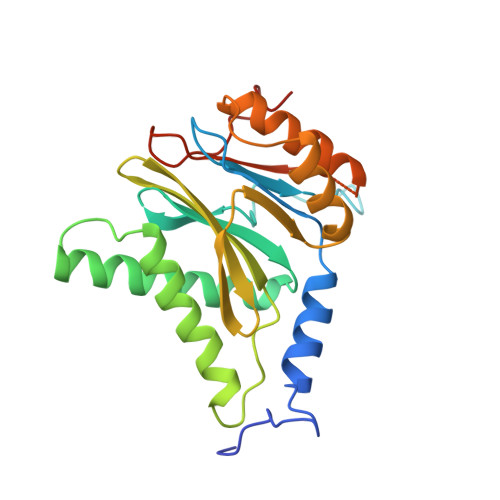
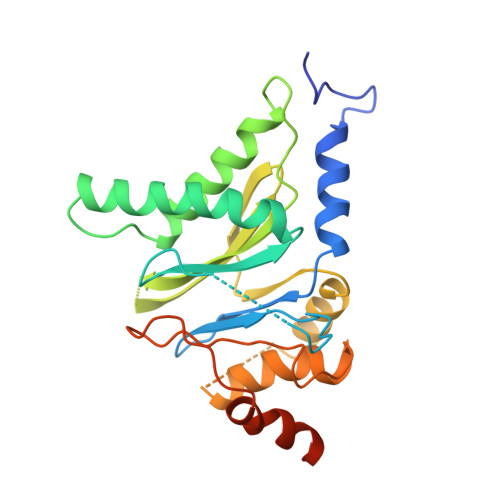
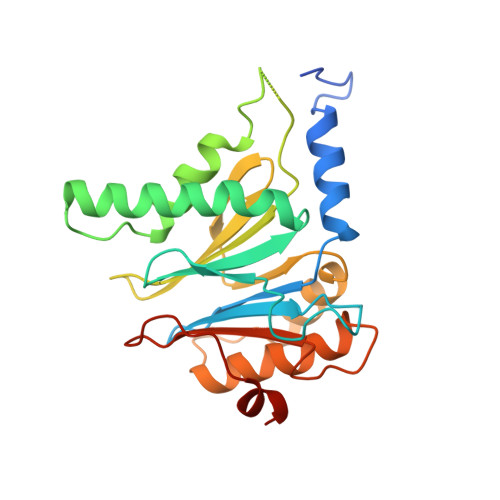
Cryo-Em Reveals the Conformation of a Substrate Analogue in the Human 20S Proteasome Core.
Da Fonseca, P.C.A., Morris, E.P.(2015) Nat Commun 6: 7573
- PubMed: 26133119 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/ncomms8573
- Primary Citation Related Structures:
5A0Q - PubMed Abstract:
The proteasome is a highly regulated protease complex fundamental for cell homeostasis and controlled cell cycle progression. It functions by removing a wide range of specifically tagged proteins, including key cellular regulators. Here we present the structure of the human 20S proteasome core bound to a substrate analogue inhibitor molecule, determined by electron cryo-microscopy (cryo-EM) and single-particle analysis at a resolution of around 3.5 Å. Our map allows the building of protein coordinates as well as defining the location and conformation of the inhibitor at the different active sites. These results open new prospects to tackle the proteasome functional mechanisms. Moreover, they also further demonstrate that cryo-EM is emerging as a realistic approach for general structural studies of protein-ligand interactions.
- MRC Laboratory of Molecular Biology, Francis Crick Avenue, Cambridge CB2 0QH, UK.
Organizational Affiliation: