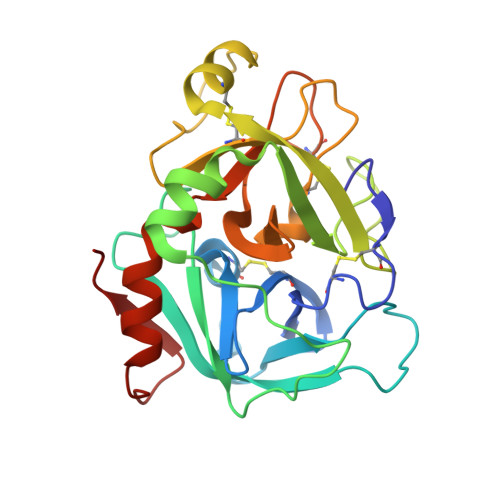
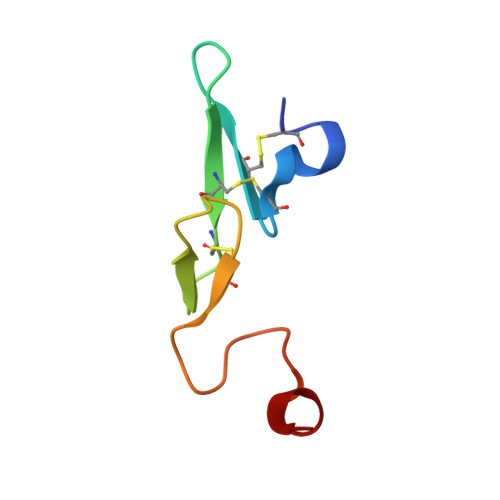
Structure-Based Design of Macrocyclic Coagulation Factor VIIa Inhibitors.
Priestley, E.S., Cheney, D.L., DeLucca, I., Wei, A., Luettgen, J.M., Rendina, A.R., Wong, P.C., Wexler, R.R.(2015) J Med Chem 58: 6225-6236
- PubMed: 26151189
- DOI: https://doi.org/10.1021/acs.jmedchem.5b00788
- Primary Citation of Related Structures:
4ZXX, 4ZXY - PubMed Abstract:
On the basis of a crystal structure of a phenylpyrrolidine lead and subsequent molecular modeling results, we designed and synthesized a novel series of macrocyclic FVIIa inhibitors. The optimal 16-membered macrocycle was 60-fold more potent than an acyclic analog. Further potency optimization by incorporation of P1' alkyl sulfone and P2 methyl groups provided a macrocycle with TF/FVIIa Ki = 1.6 nM, excellent selectivity against a panel of seven serine proteases, and FVII-deficient prothrombin time EC2x = 1.2 μM. Discovery of this potent, selective macrocyclic scaffold opens new possibilities for the development of orally bioavailable FVIIa inhibitors.