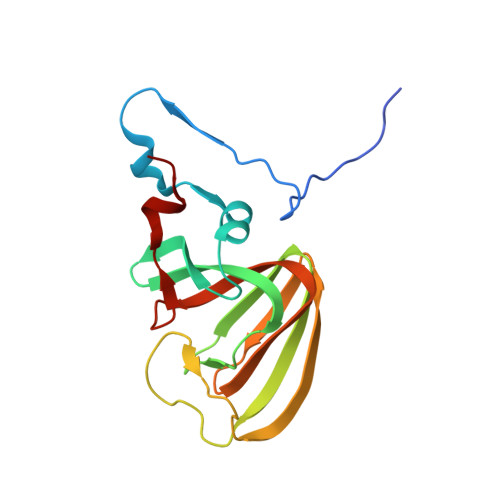
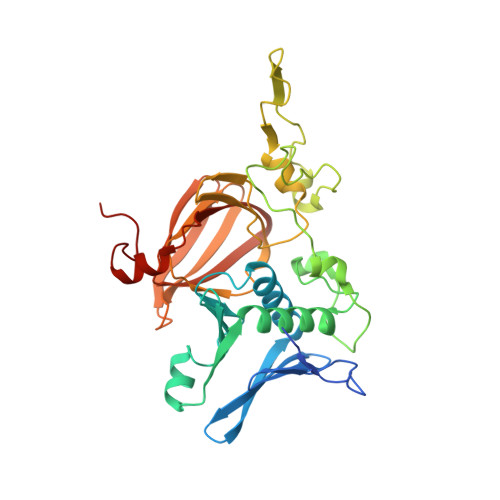
Crystal Structure of PnpCD, a Two-subunit Hydroquinone 1,2-Dioxygenase, Reveals a Novel Structural Class of Fe2+-dependent Dioxygenases.
Liu, S., Su, T., Zhang, C., Zhang, W.M., Zhu, D., Su, J., Wei, T., Wang, K., Huang, Y., Guo, L., Xu, S., Zhou, N.Y., Gu, L.(2015) J Biological Chem 290: 24547-24560
- PubMed: 26304122 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M115.673558
- Primary Citation Related Structures:
4ZXA, 4ZXC, 4ZXD - PubMed Abstract:
Aerobic microorganisms have evolved a variety of pathways to degrade aromatic and heterocyclic compounds. However, only several classes of oxygenolytic fission reaction have been identified for the critical ring cleavage dioxygenases. Among them, the most well studied dioxygenases proceed via catecholic intermediates, followed by noncatecholic hydroxy-substituted aromatic carboxylic acids. Therefore, the recently reported hydroquinone 1,2-dioxygenases add to the diversity of ring cleavage reactions. Two-subunit hydroquinone 1,2-dioxygenase PnpCD, the key enzyme in the hydroquinone pathway of para-nitrophenol degradation, catalyzes the ring cleavage of hydroquinone to γ-hydroxymuconic semialdehyde. Here, we report three PnpCD structures, named apo-PnpCD, PnpCD-Fe(3+), and PnpCD-Cd(2+)-HBN (substrate analog hydroxyenzonitrile), respectively. Structural analysis showed that both the PnpC and the C-terminal domains of PnpD comprise a conserved cupin fold, whereas PnpC cannot form a competent metal binding pocket as can PnpD cupin. Four residues of PnpD (His-256, Asn-258, Glu-262, and His-303) were observed to coordinate the iron ion. The Asn-258 coordination is particularly interesting because this coordinating residue has never been observed in the homologous cupin structures of PnpCD. Asn-258 is proposed to play a pivotal role in binding the iron prior to the enzymatic reaction, but it might lose coordination to the iron when the reaction begins. PnpD also consists of an intriguing N-terminal domain that might have functions other than nucleic acid binding in its structural homologs. In summary, PnpCD has no apparent evolutionary relationship with other iron-dependent dioxygenases and therefore defines a new structural class. The study of PnpCD might add to the understanding of the ring cleavage of dioxygenases.
- From the State Key Laboratory of Microbial Technology, School of Life Sciences, Shandong University, Jinan, Shandong 250100.
Organizational Affiliation: