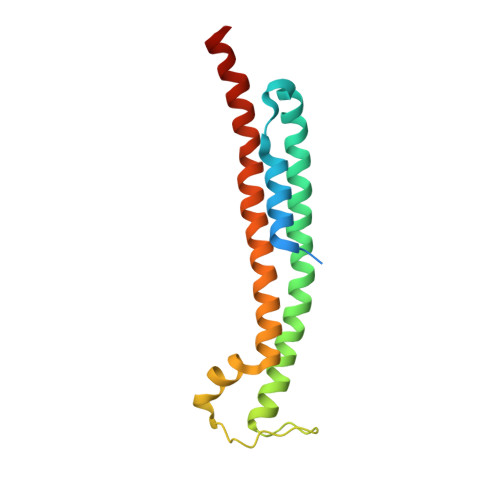
Structure and mechanism of the phage T4 recombination mediator protein UvsY.
Gajewski, S., Waddell, M.B., Vaithiyalingam, S., Nourse, A., Li, Z., Woetzel, N., Alexander, N., Meiler, J., White, S.W.(2016) Proc Natl Acad Sci U S A 113: 3275-3280
- PubMed: 26951671
- DOI: https://doi.org/10.1073/pnas.1519154113
- Primary Citation of Related Structures:
4ZWQ, 4ZWR, 4ZWS, 4ZWT - PubMed Abstract:
The UvsY recombination mediator protein is critical for efficient homologous recombination in bacteriophage T4 and is the functional analog of the eukaryotic Rad52 protein. During T4 homologous recombination, the UvsX recombinase has to compete with the prebound gp32 single-stranded binding protein for DNA-binding sites and UvsY stimulates this filament nucleation event. We report here the crystal structure of UvsY in four similar open-barrel heptameric assemblies and provide structural and biophysical insights into its function. The UvsY heptamer was confirmed in solution by centrifugation and light scattering, and thermodynamic analyses revealed that the UvsY-ssDNA interaction occurs within the assembly via two distinct binding modes. Using surface plasmon resonance, we also examined the binding of UvsY to both ssDNA and the ssDNA-gp32 complex. These analyses confirmed that ssDNA can bind UvsY and gp32 independently and also as a ternary complex. They also showed that residues located on the rim of the heptamer are required for optimal binding to ssDNA, thus identifying the putative ssDNA-binding surface. We propose a model in which UvsY promotes a helical ssDNA conformation that disfavors the binding of gp32 and initiates the assembly of the ssDNA-UvsX filament.
- Department of Structural Biology, St. Jude Children's Research Hospital, Memphis, TN 38105;
Organizational Affiliation: