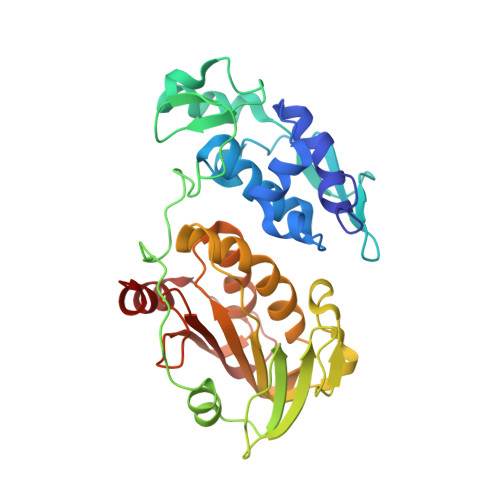
Crystal structure of chikungunya virus nsP2 cysteine protease reveals a putative flexible loop blocking its active site.
Narwal, M., Singh, H., Pratap, S., Malik, A., Kuhn, R.J., Kumar, P., Tomar, S.(2018) Int J Biol Macromol 116: 451-462
- PubMed: 29730006
- DOI: https://doi.org/10.1016/j.ijbiomac.2018.05.007
- Primary Citation Related Structures:
4ZTB - PubMed Abstract:
Chikungunya virus (CHIKV), a mosquito-borne pathogenic alphavirus is a growing public health threat. No vaccines or antiviral drug is currently available in the market for chikungunya treatment. nsP2pro, the viral cysteine protease, carries out an essential function of nonstructural polyprotein processing and forms four nonstructural proteins (nsPs) that makes the replication complex, hence constitute a promising drug target. In this study, crystal structure of nsP2pro has been determined at 2.59 Å, which reveals that the protein consists of two subdomains: an N-terminal protease subdomain and a C-terminal methyltransferase subdomain. Structural comparison of CHIKV nsP2pro with structures of other alphavirus nsP2 advances that the substrate binding cleft is present at the interface of two subdomains. Additionally, structure insights revealed that access to the active site and substrate binding cleft is blocked by a flexible interdomain loop in CHIKV nsP2pro. This loop contains His548, the catalytic residue, and Trp549 and Asn547, the residues predicted to bind substrate. Interestingly, mutation of Asn547 leads to three-fold increase in K m confirming that Asn547 plays important role in substrate binding and recognition. This study presents the detailed molecular analysis and signifies the substrate specificity residues of CHIKV nsP2pro, which will be beneficial for structure-based drug design and optimization of CHIKV protease inhibitors.
- Department of Biotechnology, Indian Institute of Technology Roorkee, Roorkee, 247667, Uttarakhand, India.
Organizational Affiliation: