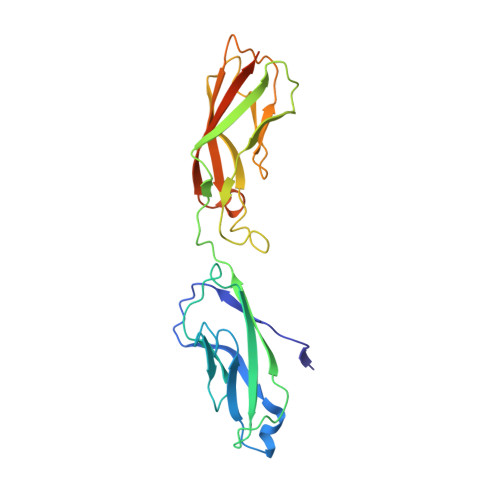
Adhesive Dimerization of Human P-Cadherin Catalyzed by a Chaperone-like Mechanism
Kudo, S., Caaveiro, J.M., Tsumoto, K.(2016) Structure 24: 1523-1536
- PubMed: 27545624
- DOI: https://doi.org/10.1016/j.str.2016.07.002
- Primary Citation Related Structures:
4ZML, 4ZMN, 4ZMO, 4ZMP, 4ZMQ, 4ZMT, 4ZMV, 4ZMW, 4ZMX, 4ZMY, 4ZMZ - PubMed Abstract:
Orderly assembly of classical cadherins governs cell adhesion and tissue maintenance. A key event is the strand-swap dimerization of the extracellular ectodomains of two cadherin molecules from apposing cells. Here we have determined crystal structures of P-cadherin in six different conformational states to elaborate a motion picture of its adhesive dimerization at the atomic level. The snapshots revealed that cell-adhesive dimerization is facilitated by several intermediate states collectively termed X-dimer in analogy to other classical cadherins. Based on previous studies and on the combined structural, kinetic, thermodynamic, biochemical, and cellular data reported herein, we propose that the adhesive dimerization of human P-cadherin is achieved by a stepwise mechanism analogous to that of assembly chaperones. This mechanism, applicable to type I classical cadherins, confers high specificity and fast association rates. We expect these findings to guide innovative therapeutic approaches targeting P-cadherin in cancer.
- Department of Chemistry & Biotechnology, The University of Tokyo, Tokyo 108-8639, Japan.
Organizational Affiliation: