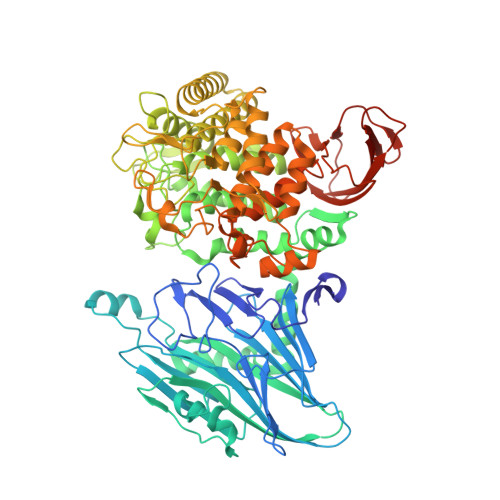
Crystal Structure and Substrate Recognition of Cellobionic Acid Phosphorylase, Which Plays a Key Role in Oxidative Cellulose Degradation by Microbes.
Nam, Y.W., Nihira, T., Arakawa, T., Saito, Y., Kitaoka, M., Nakai, H., Fushinobu, S.(2015) J Biological Chem 290: 18281-18292
- PubMed: 26041776 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M115.664664
- Primary Citation Related Structures:
4ZLE, 4ZLF, 4ZLG, 4ZLI - PubMed Abstract:
The microbial oxidative cellulose degradation system is attracting significant research attention after the recent discovery of lytic polysaccharide mono-oxygenases. A primary product of the oxidative and hydrolytic cellulose degradation system is cellobionic acid (CbA), the aldonic acid form of cellobiose. We previously demonstrated that the intracellular enzyme belonging to glycoside hydrolase family 94 from cellulolytic fungus and bacterium is cellobionic acid phosphorylase (CBAP), which catalyzes reversible phosphorolysis of CbA into glucose 1-phosphate and gluconic acid (GlcA). In this report, we describe the biochemical characterization and the three-dimensional structure of CBAP from the marine cellulolytic bacterium Saccharophagus degradans. Structures of ligand-free and complex forms with CbA, GlcA, and a synthetic disaccharide product from glucuronic acid were determined at resolutions of up to 1.6 Å. The active site is located near the dimer interface. At subsite +1, the carboxylate group of GlcA and CbA is recognized by Arg-609 and Lys-613. Additionally, one residue from the neighboring protomer (Gln-190) is involved in the carboxylate recognition of GlcA. A mutational analysis indicated that these residues are critical for the binding and catalysis of the aldonic and uronic acid acceptors GlcA and glucuronic acid. Structural and sequence comparisons with other glycoside hydrolase family 94 phosphorylases revealed that CBAPs have a unique subsite +1 with a distinct amino acid residue conservation pattern at this site. This study provides molecular insight into the energetically efficient metabolic pathway of oxidized sugars that links the oxidative cellulolytic pathway to the glycolytic and pentose phosphate pathways in cellulolytic microbes.
- From the Department of Biotechnology, The University of Tokyo, 1-1-1 Yayoi, Bunkyo-ku, Tokyo 113-8657, Japan.
Organizational Affiliation: