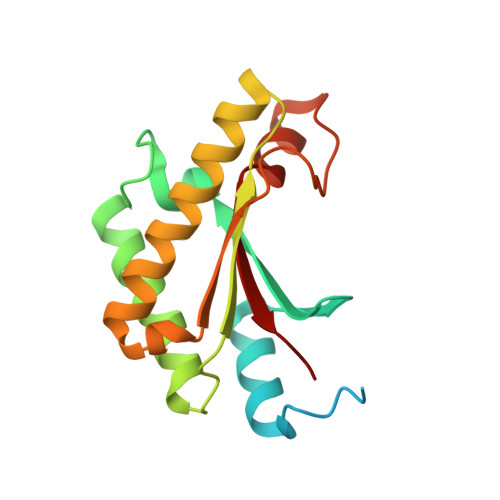
Structural insights into YfiR sequestering by YfiB in Pseudomonas aeruginosa PAO1
Li, S., Li, T., Xu, Y., Zhang, Q., Zhang, W., Che, S., Liu, R., Wang, Y., Bartlam, M.(2015) Sci Rep 5: 16915-16915
- PubMed: 26593397 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/srep16915
- Primary Citation Related Structures:
4ZHU, 4ZHV, 4ZHW, 4ZHY - PubMed Abstract:
YfiBNR is a tripartite signalling system in Pseudomonas aeruginosa that modulates intracellular c-di-GMP levels in response to signals received in the periplasm. YfiB is an outer membrane lipoprotein and presumed sensor protein that sequesters the repressor protein YfiR. To provide insights into YfiBNR function, we have determined three-dimensional crystal structures of YfiB and YfiR from P. aeruginosa PAO1 alone and as a 1:1 complex. A YfiB(27-168) construct is predominantly dimeric, whereas a YfiB(59-168) is monomeric, indicating that YfiB can dimerize via its N-terminal region. YfiR forms a stable complex with YfiB(59-168), while the YfiR binding interface is obstructed by the N-terminal region in YfiB(27-168). The YfiB-YfiR complex reveals a conserved interaction surface on YfiR that overlaps with residues predicted to interact with the periplasmic PAS domain of YfiN. Comparison of native and YfiR-bound structures of YfiB suggests unwinding of the N-terminal linker region for attachment to the outer membrane. A model is thus proposed for YfiR sequestration at the outer membrane by YfiB. Our work provides the first detailed insights into the interaction between YfiB and YfiR at the molecular level and is a valuable starting point for further functional and mechanistic studies of the YfiBNR signalling system.
- State Key Laboratory of Medicinal Chemical Biology, Nankai University, Tianjin, China.
Organizational Affiliation: