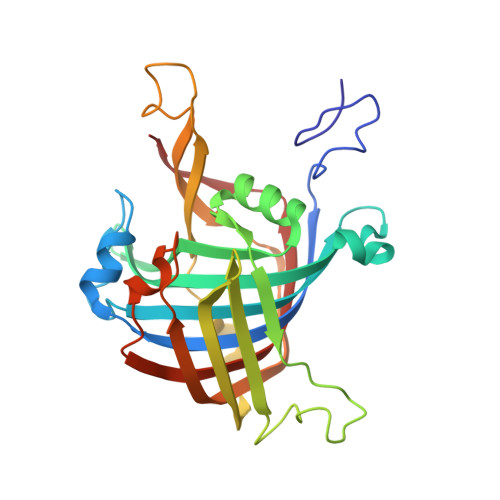
Sbi00515, a Protein of Unknown Function from Streptomyces bingchenggensis, Highlights the Functional Versatility of the Acetoacetate Decarboxylase Scaffold.
Mydy, L.S., Hoppe, R.W., Ochsenwald, J.M., Berndt, R.T., Severin, G.B., Schwabacher, A.W., Silvaggi, N.R.(2015) Biochemistry 54: 3978-3988
- PubMed: 26039798
- DOI: https://doi.org/10.1021/acs.biochem.5b00483
- Primary Citation Related Structures:
4ZBO, 4ZBT - PubMed Abstract:
The acetoacetate decarboxylase-like superfamily (ADCSF) is a group of ~4000 enzymes that, until recently, was thought to be homogeneous in terms of the reaction catalyzed. Bioinformatic analysis shows that the ADCSF consists of up to seven families that differ primarily in their active site architectures. The soil-dwelling bacterium Streptomyces bingchenggensis BCW-1 produces an ADCSF enzyme of unknown function that shares a low level of sequence identity (~20%) with known acetoacetate decarboxylases (ADCs). This enzyme, Sbi00515, belongs to the MppR-like family of the ADCSF because of its similarity to the mannopeptimycin biosynthetic protein MppR from Streptomyces hygroscopicus. Herein, we present steady state kinetic data that show Sbi00515 does not catalyze the decarboxylation of any α- or β-keto acid tested. Rather, we show that Sbi00515 catalyzes the condensation of pyruvate with a number of aldehydes, followed by dehydration of the presumed aldol intermediate. Thus, Sbi00515 is a pyruvate aldolase-dehydratase and not an acetoacetate decarboxylase. We have also determined the X-ray crystal structures of Sbi00515 in complexes with formate and pyruvate. The structures show that the overall fold of Sbi00515 is nearly identical to those of both ADC and MppR. The pyruvate complex is trapped as the Schiff base, providing evidence that the Schiff base chemistry that drives the acetoacetate decarboxylases has been co-opted to perform a new function, and that this core chemistry may be conserved across the superfamily. The structures also suggest possible catalytic roles for several active site residues.
- Department of Chemistry and Biochemistry, University of Wisconsin-Milwaukee, 3210 North Cramer Street, Milwaukee, Wisconsin 53211, United States.
Organizational Affiliation: