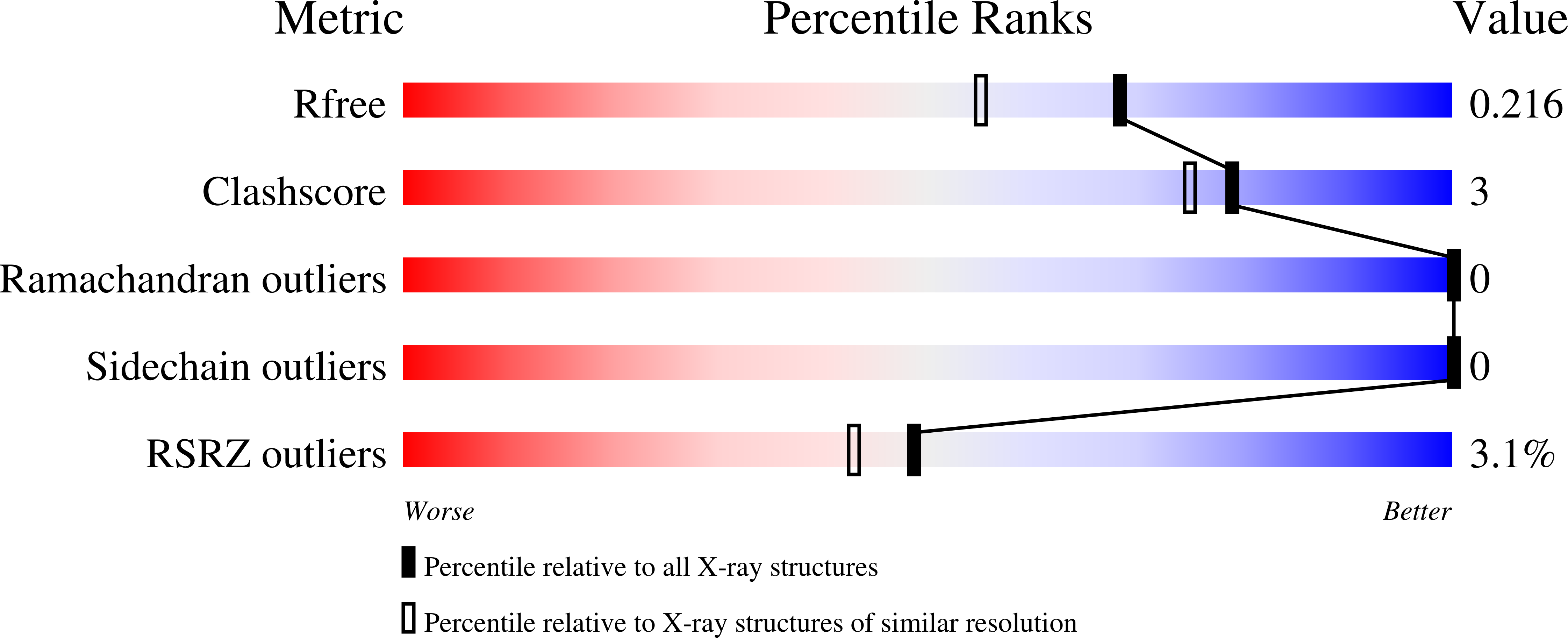
Structural studies of the yeast DNA damage-inducible protein Ddi1 reveal domain architecture of this eukaryotic protein family.
Trempe, J.F., Saskova, K.G., Siva, M., Ratcliffe, C.D., Veverka, V., Hoegl, A., Menade, M., Feng, X., Shenker, S., Svoboda, M., Kozisek, M., Konvalinka, J., Gehring, K.(2016) Sci Rep 6: 33671-33671
- PubMed: 27646017 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/srep33671
- Primary Citation Related Structures:
2N7E, 4Z2Z, 5KES - PubMed Abstract:
The eukaryotic Ddi1 family is defined by a conserved retroviral aspartyl protease-like (RVP) domain found in association with a ubiquitin-like (UBL) domain. Ddi1 from Saccharomyces cerevisiae additionally contains a ubiquitin-associated (UBA) domain. The substrate specificity and role of the protease domain in the biological functions of the Ddi family remain unclear. Yeast Ddi1 has been implicated in the regulation of cell cycle progression, DNA-damage repair, and exocytosis. Here, we investigated the multi-domain structure of yeast Ddi1 using X-ray crystallography, nuclear magnetic resonance, and small-angle X-ray scattering. The crystal structure of the RVP domain sheds light on a putative substrate recognition site involving a conserved loop. Isothermal titration calorimetry confirms that both UBL and UBA domains bind ubiquitin, and that Ddi1 binds K48-linked diubiquitin with enhanced affinity. The solution NMR structure of a helical domain that precedes the protease displays tertiary structure similarity to DNA-binding domains from transcription regulators. Our structural studies suggest that the helical domain could serve as a landing platform for substrates in conjunction with attached ubiquitin chains binding to the UBL and UBA domains.
- Groupe de Recherche Axé sur la Structure des Protéines, Department of Biochemistry, McGill University, 3649 Promenade Sir William Osler, Montreal, QC, H3G 0B1, Canada.
Organizational Affiliation: