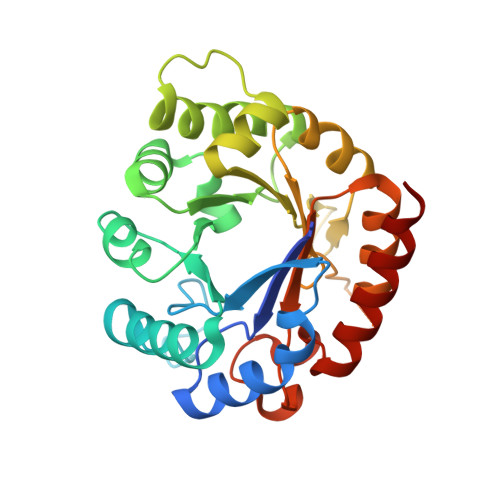
Identification of novel scaffolds for potential anti-Helicobacter pylori agents based on the crystal structure of H. pylori 3-deoxy-d-manno-octulosonate 8-phosphate synthase (HpKDO8PS).
Cho, S., Im, H., Lee, K.Y., Chen, J., Kang, H.J., Yoon, H.J., Min, K.H., Lee, K.R., Park, H.J., Lee, B.J.(2016) Eur J Med Chem 108: 188-202
- PubMed: 26649906
- DOI: https://doi.org/10.1016/j.ejmech.2015.11.036
- Primary Citation Related Structures:
4Z1A, 4Z1B, 4Z1C, 4Z1D - PubMed Abstract:
The crystal structure of 3-deoxy-d-manno-octulosonate-8-phosphate synthase (KDO8PS) from Helicobacter pylori (HpKDO8PS) was determined alone and within various complexes, revealing an extra helix (HE) that is absent in the structures of KDO8PS from other organisms. In contrast to the metal coordination of the KDO8PS enzyme from Aquifex aeolicus, HpKDO8PS is specifically coordinated with Cd(2+) or Zn(2+) ions, and isothermal titration calorimetry (ITC) and differential scanning fluorimetry (DSF) revealed that Cd(2+) thermally stabilizes the protein structure more efficiently than Zn(2+). In the substrate-bound structure, water molecules play a key role in fixing residues in the proper configuration to achieve a compact structure. Using the structures of HpKDO8PS and API [arabinose 5-phosphate (A5P) and phosphoenolpyruvate (PEP) bisubstrate inhibitor], we generated 21 compounds showing potential HpKDO8PS-binding properties via in silico virtual screening. The capacity of three, avicularin, hyperin, and MC181, to bind to HpKDO8PS was confirmed through saturation transfer difference (STD) experiments, and we identified their specific ligand binding modes by combining competition experiments and docking simulation analysis. Hyperin was confirmed to bind to the A5P binding site, primarily via hydrophilic interaction, whereas MC181 bound to both the PEP and A5P binding sites through hydrophilic and hydrophobic interactions. These results were consistent with the epitope mapping by STD. Our results are expected to provide clues for the development of HpKDO8PS inhibitors.
- Research Institute of Pharmaceutical Sciences, College of Pharmacy, Seoul National University, Seoul 151-742, Republic of Korea.
Organizational Affiliation: