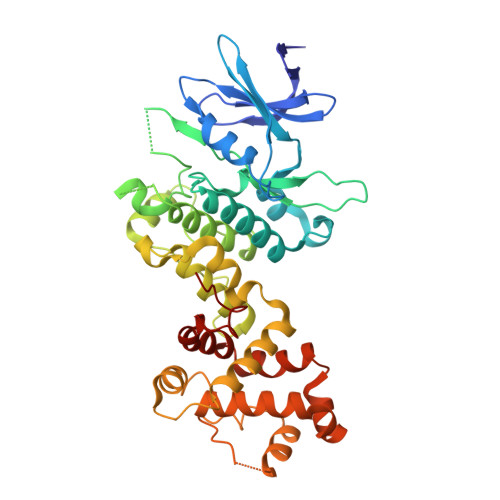
Long-Range Inhibitor-Induced Conformational Regulation of Human IRE1 alpha Endoribonuclease Activity.
Concha, N.O., Smallwood, A., Bonnette, W., Totoritis, R., Zhang, G., Federowicz, K., Yang, J., Qi, H., Chen, S., Campobasso, N., Choudhry, A.E., Shuster, L.E., Evans, K.A., Ralph, J., Sweitzer, S., Heerding, D.A., Buser, C.A., Su, D.S., DeYoung, M.P.(2015) Mol Pharmacol 88: 1011-1023
- PubMed: 26438213 Search on PubMed
- DOI: https://doi.org/10.1124/mol.115.100917
- Primary Citation Related Structures:
4YZ9, 4YZC, 4YZD - PubMed Abstract:
Activation of the inositol-requiring enzyme-1 alpha (IRE1α) protein caused by endoplasmic reticulum stress results in the homodimerization of the N-terminal endoplasmic reticulum luminal domains, autophosphorylation of the cytoplasmic kinase domains, and conformational changes to the cytoplasmic endoribonuclease (RNase) domains, which render them functional and can lead to the splicing of X-box binding protein 1 (XBP 1) mRNA. Herein, we report the first crystal structures of the cytoplasmic portion of a human phosphorylated IRE1α dimer in complex with (R)-2-(3,4-dichlorobenzyl)-N-(4-methylbenzyl)-2,7-diazaspiro(4.5)decane-7-carboxamide, a novel, IRE1α-selective kinase inhibitor, and staurosporine, a broad spectrum kinase inhibitor. (R)-2-(3,4-dichlorobenzyl)-N-(4-methylbenzyl)-2,7-diazaspiro(4.5)decane-7-carboxamide inhibits both the kinase and RNase activities of IRE1α. The inhibitor interacts with the catalytic residues Lys599 and Glu612 and displaces the kinase activation loop to the DFG-out conformation. Inactivation of IRE1α RNase activity appears to be caused by a conformational change, whereby the αC helix is displaced, resulting in the rearrangement of the kinase domain-dimer interface and a rotation of the RNase domains away from each other. In contrast, staurosporine binds at the ATP-binding site of IRE1α, resulting in a dimer consistent with RNase active yeast Ire1 dimers. Activation of IRE1α RNase activity appears to be promoted by a network of hydrogen bond interactions between highly conserved residues across the RNase dimer interface that place key catalytic residues poised for reaction. These data implicate that the intermolecular interactions between conserved residues in the RNase domain are required for activity, and that the disruption of these interactions can be achieved pharmacologically by small molecule kinase domain inhibitors.
- Oncology R&D (K.F., J.Y., L.E.S., K.A.E., J.R., D.A.H., C.A.B., D.S.S, M.P.D.), Biological Sciences (R.T., G.Z., H.Q., S.C., A.E.C., S.S.), and Chemical Sciences, GlaxoSmithKline Research and Development, Collegeville, Pennsylvania (N.O.C., A.S., W.B., N.C.) maurice.p.deyoung@gsk.com Nestor.O.Concha@gsk.com.
Organizational Affiliation: