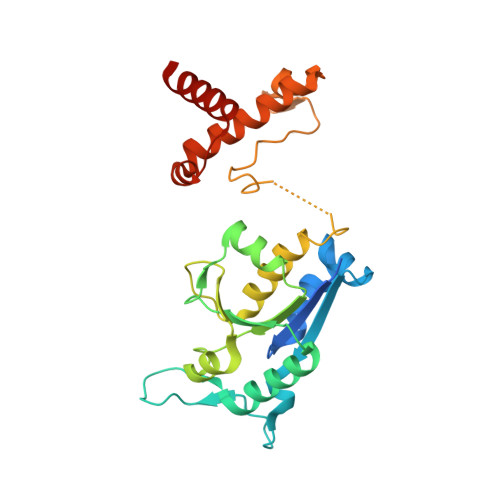
Structural basis for methyl-donor-dependent and sequence-specific binding to tRNA substrates by knotted methyltransferase TrmD.
Ito, T., Masuda, I., Yoshida, K., Goto-Ito, S., Sekine, S., Suh, S.W., Hou, Y.M., Yokoyama, S.(2015) Proc Natl Acad Sci U S A 112: E4197-E4205
- PubMed: 26183229
- DOI: https://doi.org/10.1073/pnas.1422981112
- Primary Citation Related Structures:
4YVG, 4YVH, 4YVI, 4YVJ, 4YVK - PubMed Abstract:
The deep trefoil knot architecture is unique to the SpoU and tRNA methyltransferase D (TrmD) (SPOUT) family of methyltransferases (MTases) in all three domains of life. In bacteria, TrmD catalyzes the N(1)-methylguanosine (m(1)G) modification at position 37 in transfer RNAs (tRNAs) with the (36)GG(37) sequence, using S-adenosyl-l-methionine (AdoMet) as the methyl donor. The m(1)G37-modified tRNA functions properly to prevent +1 frameshift errors on the ribosome. Here we report the crystal structure of the TrmD homodimer in complex with a substrate tRNA and an AdoMet analog. Our structural analysis revealed the mechanism by which TrmD binds the substrate tRNA in an AdoMet-dependent manner. The trefoil-knot center, which is structurally conserved among SPOUT MTases, accommodates the adenosine moiety of AdoMet by loosening/retightening of the knot. The TrmD-specific regions surrounding the trefoil knot recognize the methionine moiety of AdoMet, and thereby establish the entire TrmD structure for global interactions with tRNA and sequential and specific accommodations of G37 and G36, resulting in the synthesis of m(1)G37-tRNA.
- RIKEN Systems and Structural Biology Center, Tsurumi-ku, Yokohama 230-0045, Japan; Graduate School of Science, The University of Tokyo, Bunkyo-ku, Tokyo 113-0033, Japan; Division of Structural and Synthetic Biology, RIKEN Center for Life Science Technologies, Tsurumi-ku, Yokohama 230-0045, Japan;
Organizational Affiliation: