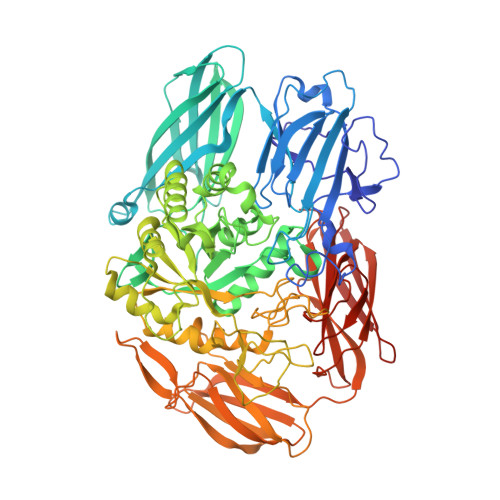
Crystal structure of beta-galactosidase from Bacillus circulans ATCC 31382 (BgaD) and the construction of the thermophilic mutants.
Ishikawa, K., Kataoka, M., Yanamoto, T., Nakabayashi, M., Watanabe, M., Ishihara, S., Yamaguchi, S.(2015) FEBS J 282: 2540-2552
- PubMed: 25879162 Search on PubMed
- DOI: https://doi.org/10.1111/febs.13298
- Primary Citation Related Structures:
4YPJ - PubMed Abstract:
β-Galactosidase (EC 3.2.1.23) from Bacillus circulans ATCC 31382, designated BgaD, exhibits high transglycosylation activity to produce galacto-oligosaccharides. BgaD has been speculated to have a multiple domain architecture including a F5/8-type C domain or a discoidin domain in the C-terminal peptide region from amino acid sequence analysis. Here, we solved the first crystal structure of the C-terminal deletion mutant BgaD-D, consisting of sugar binding, Glyco_hydro, catalytic and bacterial Ig-like domains, at 2.5 Å. In the asymmetric unit, two molecules of BgaD-D were identified and the value of VM was estimated to be 5.0 Å(3) · Da(-1). It has been speculated that BgaD-D consists of four domains. From the structural analysis, however, we clarified that BgaD-D consists of five domains. We identified a new domain structure comprised of β-sheets in BgaD. The catalytic domain exhibits a TIM barrel structure with a small pocket suited for accommodating the disaccharides. Detailed structural information for the amino acid residues related to activity and substrate specificity was clarified in the catalytic domain. Furthermore, using the structural information, we successfully constructed some thermostable mutants via protein engineering method. Coordinates for the BgaD-D structure have been deposited in the Protein Data Bank under accession code 4YPJ.
- Biomass Refinery Research Center, National Institute of Advanced Industrial Science and Technology (AIST), Higashi-hiroshima, Japan.
Organizational Affiliation: