Crystal Structure of E2 Complex
Brown, N.G., Cho, S.E., Schulman, B.A.(2015) Proc Natl Acad Sci U S A
Experimental Data Snapshot
Starting Models: experimental
View more details
wwPDB Validation 3D Report Full Report
(2015) Proc Natl Acad Sci U S A
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Ubiquitin-conjugating enzyme E2 C | A [auth U] | 155 | Homo sapiens | Mutation(s): 0 Gene Names: UBE2C, UBCH10 EC: 6.3.2.19 (PDB Primary Data), 2.3.2.24 (UniProt), 2.3.2.23 (UniProt) | 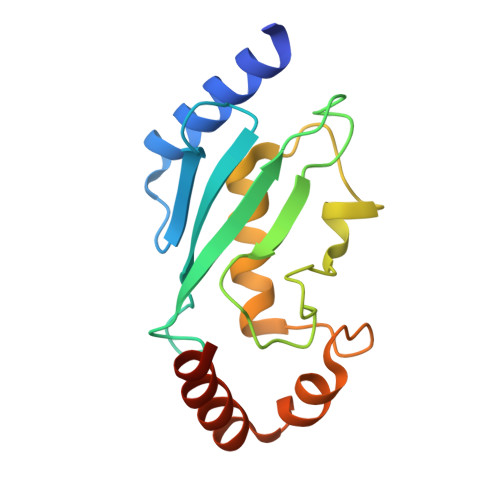 |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for O00762 (Homo sapiens) Explore O00762 Go to UniProtKB: O00762 | |||||
PHAROS: O00762 GTEx: ENSG00000175063 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | O00762 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Anaphase-promoting complex subunit 2 | B [auth A] | 90 | Homo sapiens | Mutation(s): 0 Gene Names: ANAPC2, APC2, KIAA1406 | 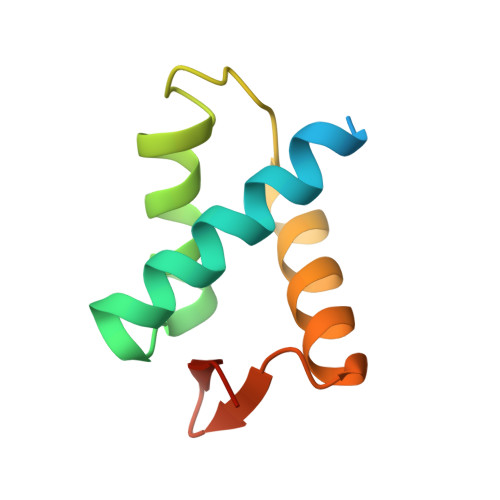 |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for Q9UJX6 (Homo sapiens) Explore Q9UJX6 Go to UniProtKB: Q9UJX6 | |||||
PHAROS: Q9UJX6 GTEx: ENSG00000176248 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q9UJX6 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 133.113 | α = 90 |
| b = 33.188 | β = 100.12 |
| c = 51.963 | γ = 90 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| HKL-2000 | data reduction |
| HKL-2000 | data scaling |
| PHASER | phasing |