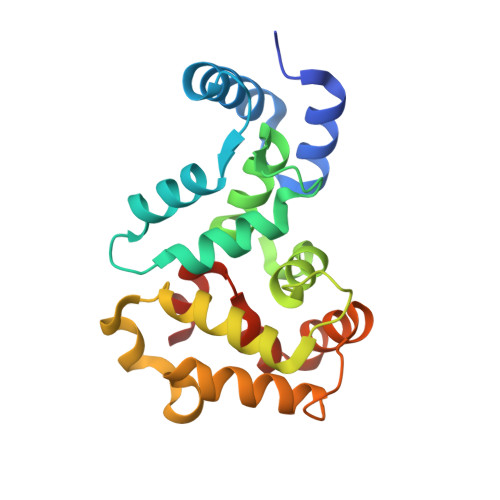
Crystal Structure of Recoverin with Calcium Ions Bound to Both Functional EF Hands.
Kumar, R.P., Ranaghan, M.J., Ganjei, A.Y., Oprian, D.D.(2015) Biochemistry 54: 7222-7228
- PubMed: 26584024 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.biochem.5b01160
- Primary Citation Related Structures:
4YI8, 4YI9 - PubMed Abstract:
Recoverin (Rv), a small Ca(2+)-binding protein that inhibits rhodopsin kinase (RK), has four EF hands, two of which are functional (EF2 and EF3). Activation requires Ca(2+) in both EF hands, but crystal structures have never been observed with Ca(2+) ions in both sites; all previous structures have Ca(2+) bound to only EF3. We suspected that this was due to an intermolecular crystal contact between T80 and a surface glutamate (E153) that precluded coordination of a Ca(2+) ion in EF2. We constructed the E153A mutant, determined its X-ray crystal structure to 1.2 Å resolution, and showed that two Ca(2+) ions are bound, one in EF3 and one in EF2. Additionally, several other residues are shown to adopt conformations in the 2Ca(2+) structure not seen previously and not seen in a second structure of the E153A mutant containing Na(+) instead of Ca(2+) in the EF2 site. The side-chain rearrangements in these residues form a 28 Å allosteric cascade along the surface of the protein connecting the Ca(2+)-binding site of EF2 with the active-site pocket responsible for binding RK.
- Department of Biochemistry, Brandeis University , Waltham, Massachusetts 02454, United States.
Organizational Affiliation: