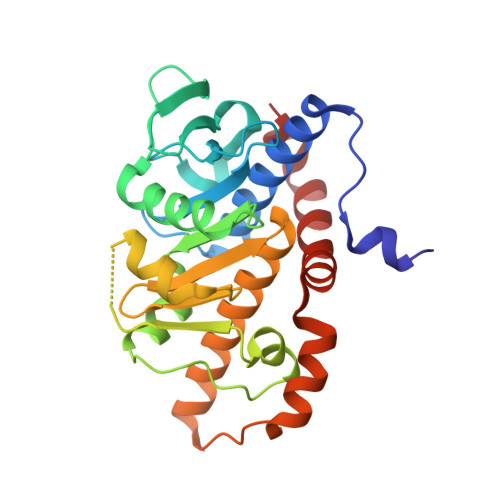
Crystal structure of fuculose aldolase from the Antarctic psychrophilic yeast Glaciozyma antarctica PI12.
Jaafar, N.R., Littler, D., Beddoe, T., Rossjohn, J., Illias, R.M., Mahadi, N.M., Mackeen, M.M., Murad, A.M., Abu Bakar, F.D.(2016) Acta Crystallogr F Struct Biol Commun 72: 831-839
- PubMed: 27827354 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X16015612
- Primary Citation Related Structures:
4XXF - PubMed Abstract:
Fuculose-1-phosphate aldolase (FucA) catalyses the reversible cleavage of L-fuculose 1-phosphate to dihydroxyacetone phosphate (DHAP) and L-lactaldehyde. This enzyme from mesophiles and thermophiles has been extensively studied; however, there is no report on this enzyme from a psychrophile. In this study, the gene encoding FucA from Glaciozyma antarctica PI12 (GaFucA) was cloned and the enzyme was overexpressed in Escherichia coli, purified and crystallized. The tetrameric structure of GaFucA was determined to 1.34 Å resolution. The overall architecture of GaFucA and its catalytically essential histidine triad are highly conserved among other fuculose aldolases. Comparisons of structural features between GaFucA and its mesophilic and thermophilic homologues revealed that the enzyme has typical psychrophilic attributes, indicated by the presence of a high number of nonpolar residues at the surface and a lower number of arginine residues.
- School of Biosciences and Biotechnology, Faculty of Science and Technology, Universiti Kebangsaan Malaysia, Bangi, Selangor Darul Ehsan 43600, Malaysia.
Organizational Affiliation: