Discovery of N-(4-aryl-5-aryloxy-thiazol-2-yl)-amides as potent ROR gamma t inverse agonists
Wang, Y., Yang, T., Liu, Q., Ma, Y., Yang, L., Zhou, L., Xiang, Z., Cheng, Z., Lu, S., Orband-Miller, L.A., Zhang, W., Wu, Q., Zhang, K., Li, Y., Xiang, J.N., Elliott, J.D., Leung, S., Ren, F., Lin, X.(2015) Bioorg Med Chem 23: 5293-5302
- PubMed: 26277758 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmc.2015.07.068
- Primary Citation Related Structures:
4XT9 - PubMed Abstract:
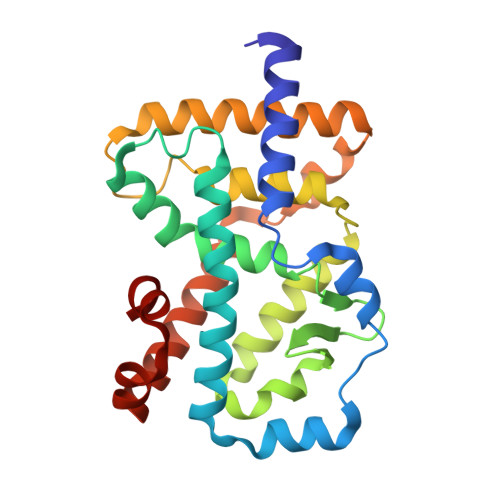
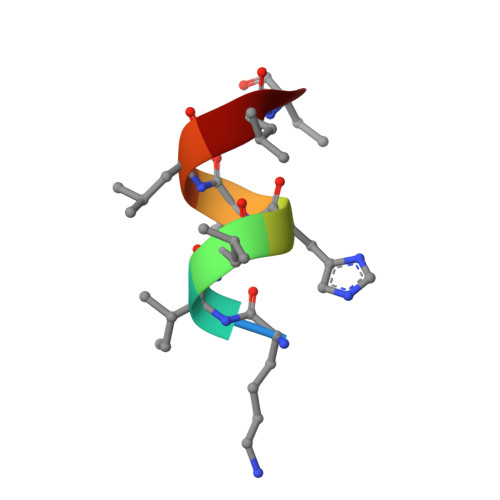
A novel series of N-(4-aryl-5-aryloxy-thiazol-2-yl)-amides as RORγt inverse agonists was discovered. Binding mode analysis of a RORγt partial agonist (2c) revealed by co-crystal structure in RORγt LBD suggests that the inverse agonists do not directly interfere with the interaction between H12 and the RORγt LBD. Detailed SAR exploration led to identification of potent RORγt inverse agonists such as 3m with a pIC50 of 8.0. Selected compounds in the series showed reasonable activity in Th17 cell differentiation assay as well as low intrinsic clearance in mouse liver microsomes.
- School of Pharmacy, Fudan University, 826 Zhangheng Road, Pudong, Shanghai 201203, China. Electronic address: yonghuiwang@fudan.edu.cn.
Organizational Affiliation: