Enthalpic cost of water removal from a hydrophobic glucose binding cavity on HK620 tailspike protein.
Gohlke, U., Broeker, N.K., Kunstmann, S., Santer, M., Heinemann, U., Lipowski, R., Seckler, R., Barbirz, S.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
wwPDB Validation 3D Report Full Report
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Tail spike protein | 597 | Enterobacteria phage HK620 | Mutation(s): 2 | 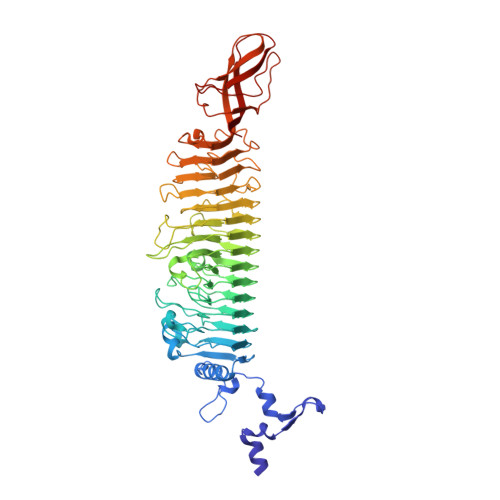 | |
UniProt | |||||
Find proteins for Q9AYY6 (Enterobacteria phage HK620) Explore Q9AYY6 Go to UniProtKB: Q9AYY6 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q9AYY6 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Length | 2D Diagram | Glycosylation | 3D Interactions |
| alpha-L-rhamnopyranose-(1-6)-alpha-D-glucopyranose-(1-4)-[2-acetamido-2-deoxy-beta-D-glucopyranose-(1-3)]alpha-D-galactopyranose-(1-3)-2-acetamido-2-deoxy-alpha-D-glucopyranose | B | 6 |  | N/A | |
Glycosylation Resources | |||||
GlyTouCan: G81480NI GlyCosmos: G81480NI | |||||
| Ligands 3 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| TRS Query on TRS | C [auth A] | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL C4 H12 N O3 LENZDBCJOHFCAS-UHFFFAOYSA-O |  | ||
| FMT Query on FMT | D [auth A], E [auth A], F [auth A], G [auth A] | FORMIC ACID C H2 O2 BDAGIHXWWSANSR-UHFFFAOYSA-N |  | ||
| NA Query on NA | H [auth A], I [auth A], J [auth A] | SODIUM ION Na FKNQFGJONOIPTF-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 74.257 | α = 90 |
| b = 74.257 | β = 90 |
| c = 174.512 | γ = 120 |
| Software Name | Purpose |
|---|---|
| XDS | data reduction |
| Aimless | data scaling |
| PHASER | phasing |
| ARP | model building |
| Coot | model building |
| REFMAC | refinement |
| PDB_EXTRACT | data extraction |
| Funding Organization | Location | Grant Number |
|---|---|---|
| German Research Foundation | Germany | BA 4046/1-1 |