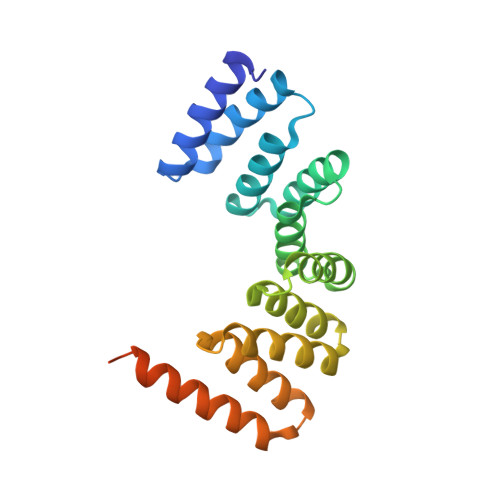
MamA as a Model Protein for Structure-Based Insight into the Evolutionary Origins of Magnetotactic Bacteria.
Zeytuni, N., Cronin, S., Lefevre, C.T., Arnoux, P., Baran, D., Shtein, Z., Davidov, G., Zarivach, R.(2015) PLoS One 10: e0130394-e0130394
- PubMed: 26114501 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0130394
- Primary Citation Related Structures:
4XI0 - PubMed Abstract:
MamA is a highly conserved protein found in magnetotactic bacteria (MTB), a diverse group of prokaryotes capable of navigating according to magnetic fields - an ability known as magnetotaxis. Questions surround the acquisition of this magnetic navigation ability; namely, whether it arose through horizontal or vertical gene transfer. Though its exact function is unknown, MamA surrounds the magnetosome, the magnetic organelle embedding a biomineralised nanoparticle and responsible for magnetotaxis. Several structures for MamA from a variety of species have been determined and show a high degree of structural similarity. By determining the structure of MamA from Desulfovibrio magneticus RS-1 using X-ray crystallography, we have opened up the structure-sequence landscape. As such, this allows us to perform structural- and phylogenetic-based analyses using a variety of previously determined MamA from a diverse range of MTB species across various phylogenetic groups. We found that MamA has remained remarkably constant throughout evolution with minimal change between different taxa despite sequence variations. These findings, coupled with the generation of phylogenetic trees using both amino acid sequences and 16S rRNA, indicate that magnetotaxis likely did not spread via horizontal gene transfer and instead has a significantly earlier, primordial origin.
- Department of Life Sciences and The National Institute for Biotechnology in the Negev, Ben-Gurion University of the Negev, Beer Sheva, Israel.
Organizational Affiliation: